Metagenomics
Launch Galaxy| Public Server: | Server (DE) |
|---|---|
| Scope: | Domain Specific |
| Summary: | a webserver to process, analyse and visualize Metagenomic and Microbiota data in general. |
Comments
- More than 200 tools are integrated in this custom Galaxy instance. They were chosen for their use in exploitation of microbiota data.
- The public server is hosted by the UseGalaxy.eu team.
Quotas
- Storage and computational quotas.
Citations
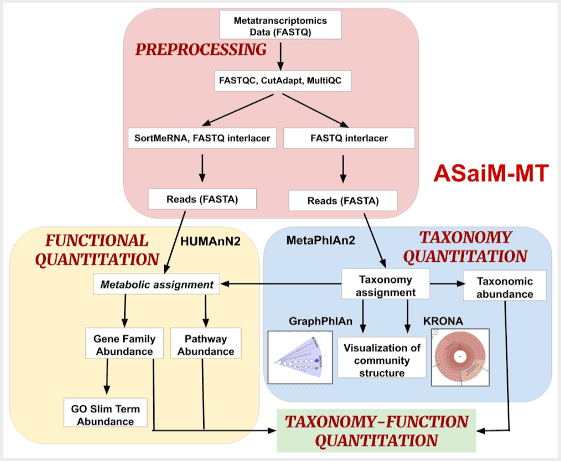
- Mehta, S., Crane, M., Leith, E., Batut, B., Hiltemann, S., Arntzen, M. Ø., Kunath, B. J., Delogu, F., Sajulga, R., Kumar, P., Johnson, J. E., Griffin, T. J., & Jagtap, P. D. (2021). ASaiM-MT: A validated and optimized ASaiM workflow for metatranscriptomics analysis within Galaxy framework. F1000Research, 10, 103. doi: 10.12688/f1000research.28608.1
- Batut, B., Gravouil, K., Defois, C., Hiltemann, S., Brugère, J.-F., Peyretaillade, E., & Peyret, P. (2018). ASaiM: A Galaxy-based framework to analyze microbiota data. GigaScience, 7(6). doi: 10.1093/gigascience/giy057
- Metagenomics EU tagged publications in the Galaxy Publication Library
Sponsors
- The Freiburg Galaxy Team but also collectively by groups and individuals from across Europe