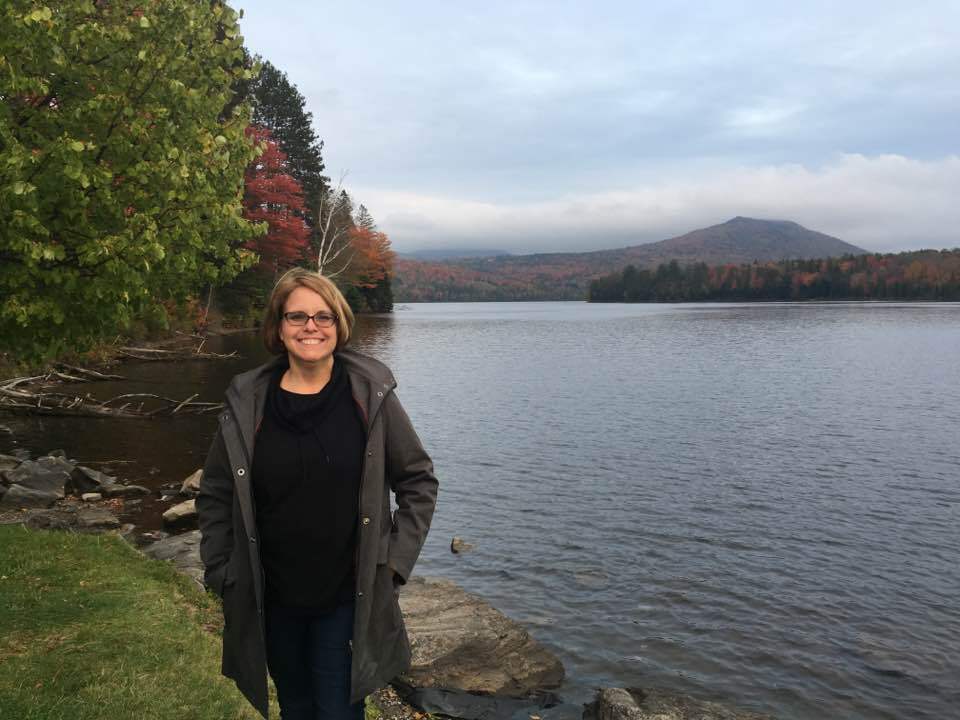
GCC2021 Training Week

In June/July 2021, we will organize a global 5-day Galaxy Training event as part of the Galaxy Community Conference (GCC2021)
This will be an online event, spanning all time zones. All training sessions are pre-recorded, so you can work through them at your own pace, and manage your own time. A large community of GTN trainers will be available 24/7 online support to answer all your questions on Slack (a chat platform).
We will have 3 Training Tracks, aimed at scientists, developers and system administrators.
The Galaxy for Scientists Track will cover a wide range of topics including NGS Analysis (DNA & RNA), Proteomics, Microbial Analysis, Machine Learning, using R in Galaxy, Visualisation, Clinical workflows (including COVID-19), Nanopore Analysis, Single-cell RNASeq, Epigenetics, Climate Science, Cheminformatics, Advanced Galaxy features (scaling your analysis) and more! YOU can choose which of these training modules you want to follow during the week of training.
The Galaxy for Developers Track will teach you how Galaxy works internally (frontend and backend), how to add new features for Galaxy, how to build the best tools and visualisations, how to interact with the Galaxy API using BioBlend, and more!
The Galaxy for Administrators Track teaches you how to be a stellar admin, easily deploying a production-ready Galaxy with Ansible!
The Science and Developer tracks do not have a predetermined path, there will be a number of training “modules” covering a variety of topics from which you can choose to build your own training program for the week. For the Admin track, there is a set 5-day curriculum that will guide you through the end-to-end process of building your own Galaxy. For all 3 tracks you can work through the materials at your own pace and according to your own time schedule. A global community of instructors will be online to provide 24/7 support to help you with these trainin sessions.
Please see the full program below for more details on the training
Practical Information
When: June 28- July 2, 2021 (all time zones)
Who: Participants of the Galaxy Community Conference
Format: Virtual and asynchronous. All training session are pre-recorded, you can work through these at your own pace, with instructors available online for support
Support: Slack and Remo
Contact: For questions regarding registration, please contact the VIB Conference Team, for any other questions, contact Saskia Hiltemann, Helena Rasche
Registration
Please sign up the Galaxy Community Conference to join this event. Registration can be done via the VIB Conference Website.
Many people have had questions about tickets, so, here is a short guide of which tickets you need for which training tracks:
- If you want to do admin training you need an admin ticket
- If you want to do either science or developer training, you need a non-admin ticket.
Or as a table to list all of the combinations
| Admin | Science | Dev | Which Tickets you need |
|---|---|---|---|
| No | No | No | None! |
| No | No | Yes | Non-admin |
| No | Yes | No | Non-admin |
| No | Yes | Yes | Non-admin |
| Yes | No | No | Admin |
| Yes | No | Yes | Admin + Non-admin |
| Yes | Yes | No | Admin + Non-admin |
| Yes | Yes | Yes | Admin + Non-admin |
Program
Welcome! We will have 3 tracks during this GCC Training Week. A short overview of the program for each of these sessions can also be found below.
Note: The schedule is subject to change
Science Track: Overview table
Welcome & Practical Information
Start here. This will cover all the logistics and practical information for this training week.
| Topic | Presentor |
|---|---|
| Welcome & Course Information | Helena Rasche, Saskia Hiltemann |
| Get set up for the course | |
| About Certificates | |
| Meet & Join the Galaxy Community! | The Global GTN Community |
| Daily Icebreaker Questions |
Introduction to Galaxy
Start here if you are new to Galaxy. These videos will introduce you to the Galaxy platform, and walk you through your first analyses
| Topic | Presentor |
|---|---|
| A Very Short Introduction to Galaxy | Anton Nekrutenko |
| Your First Galaxy Analysis (genomics-based) | Dave Clements |
| Galaxy Intro for Everyone (non-genomics) | Anne Fouilloux |
| Webinar Series: Galaxy Resources For.. |
Advanced Galaxy Features
These tutorials cover some more advanced Galaxy features
| Topic | Presentor |
|---|---|
| Uploading data with the Rule Based Uploader | Assunta DeSanto |
| Running Workflows from the Command Line using Planemo | Simon Bray |
Introduction to NGS Analysis
This module will introduce the basics of NGS analysis, from cleaning your data to mapping and assembly
| Topic | Presentor |
|---|---|
| Demo: NGS Data Logistics using SARS-CoV-2 data | Anton Nekrutenko |
| Quality Control | Florian Heyl |
| Mapping | Peter van Heusden |
| Genome Assembly |
Advanced NGS Analysis
| Topic | Presentor |
|---|---|
| Assembly: Unicycler assembly of SARS-CoV-2 genome | Cristóbal Gallardo |
| Genome Annotation: Annotation of a prokaryotic genome | |
| Genome Annotation: Refining Genome Annotations with Apollo | Anthony Bretaudeau |
| Epigenetics - ATAC-Seq data analysis | |
| SRA Aligned Read Formats to Speed Up SARS-CoV-2 Analysis | Jonathan Trow |
Transcriptomics
The following module cover transcriptomic analysis in Galaxy, from bulk RNA-seq to Single-cell RNA-Seq, from Quality control to Visualisation
Introduction to RNA-Seq
| Topic | Presentor |
|---|---|
| Introduction to RNA-Seq | Fotis E. Psomopoulos |
| Reference-based RNA-seq Analysis | Bérénice Batut |
| De novo transcriptome reconstruction with RNA-Seq | Mallory Freeberg |
| Whole transcriptome analysis of Arabidopsis thaliana | Cristóbal Gallardo |
RNA-Seq analysis using Rstudio in Galaxy
| Topic | Presentor |
|---|---|
| Starting Rstudio in Galaxy | Fotis E. Psomopoulos |
| Introduction to R | Fotis E. Psomopoulos |
| Advanced R in Galaxy | Fotis E. Psomopoulos |
| Visualisation: RNASeq postprocessing with R | Fotis E. Psomopoulos |
| Visualization of RNA-Seq results with Volcano Plot in Galaxy and R | Maria Doyle |
Single-cell RNA-Seq Analysis
Why smash tissues up to analyse them, when you can find what’s inside each individual cell? In these scRNA-seq tutorials, you’ll move from FASTQ to trajectory using the same dataset throughout – a case study of previously published, real mouse data. And for the plant enthusiasts, you’ll do the same thing in Arabidopsis! Enjoy life at a single cell scale!
| Topic | Presentor |
|---|---|
| An introduction to scRNA-seq data analysis | These slides are narrated by AWS Polly. |
| Generating a single cell matrix using Alevin | Wendi Bacon |
| Filter, Plot and Explore Single-cell RNA-seq Data | Wendi Bacon |
| Trajectory Analysis using Python (Jupyter Notebook) in Galaxy | Wendi Bacon |
| Analysis of plant scRNA-Seq Data with Scanpy | Mehmet Tekman |
Proteomics
In this module we explore the world of proteomics! Today we have a mixture of lectures, hands-on tutorials, and workflow demonstrations. The FAQ document provides links to example histories if you would like to explore the outputs of the demos yourself.
| Topic | Presentor |
|---|---|
| Introduction to mass spectrometry based proteomics data analysis | These slides are narrated by AWS Polly. |
| Maxquant + MSstats on a clinical cohort | Melanie Föll |
| DIA Mass-spectrometry with EncyclopeDIA | Emma Leith, James Johnson, Pratik Jagtap |
| ProteoRE tool for Functional Analysis of a Protein List | Yves Vandenbrouck |
| ProteoRE tool for biomarker discovery | |
| Metaproteomics Introduction | Pratik Jagtap |
| Pandemics Research using Mass Spectrometry | Timothy J. Griffin, Subina Mehta, Andrew Rajczewski, Pratik Jagtap |
Proteogenomics
Proteogenomics utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides
| Topic | Presentor |
|---|---|
| Introduction to Proteogenomics | Timothy J. Griffin |
| Proteogenomics 1: Database Creation | James Johnson |
| Proteogenomics 2: Database search | Andrew Rajczewski |
| Proteogenomics 3: Novel peptide analysis | Subina Mehta |
Microbial Analysis using Galaxy
| Topic | Presentor |
|---|---|
| Variant Analysis: M. Tuberculosis Variant Analysis | Peter van Heusden |
| 16S Microbial Analysis with mothur | Saskia Hiltemann |
| Antimicrobial Resistance Detection using Nanopore long-read sequencing | |
| Metatranscriptomics analysis using microbiome RNA-seq data | Pratik Jagtap, Timothy J. Griffin, Subina Mehta, Saskia Hiltemann |
| Metaproteomics Introduction | Pratik Jagtap |
Machine Learning
| Topic | Presentor |
|---|---|
| Introduction to Machine Learning | Anup Kumar |
| Classification in Machine Learning | Anup Kumar |
| Regression in Machine Learning | Anup Kumar |
| Machine Learning using R in Galaxy | Fotis E. Psomopoulos |
| Deep Neural Networks Demystified | Kaivan Kamali |
SARS-CoV-2 analysis
Here we have collected all the training sessions that cover SARS-CoV-2 analysis. These sessions also appear in other modules and cover a range of topics, but one thing they have in common is that they use SARS-CoV-2 or COVID-19 data.
| Topic | Presentor |
|---|---|
| Demo: NGS Data Logistics using SARS-CoV-2 data | Anton Nekrutenko |
| SRA Aligned Read Formats to Speed Up SARS-CoV-2 Analysis | Jonathan Trow |
| Assembly: Unicycler assembly of SARS-CoV-2 genome | Cristóbal Gallardo |
| Pandemics Research using Mass Spectrometry | Timothy J. Griffin, Subina Mehta, Andrew Rajczewski, Pratik Jagtap |
Plant data analysis
Here we have collected all the training sessions that cover plant data analysis. These sessions also appear in other modules and cover a range of topics, but one thing they have in common is that they use plant datasets.
| Topic | Presentor |
|---|---|
| Whole transcriptome analysis of Arabidopsis thaliana | Cristóbal Gallardo |
| Analysis of plant scRNA-Seq Data with Scanpy | Mehmet Tekman |
Visualisation in Galaxy
This module will cover various ways of visualizing your data in Galaxy
| Topic | Presentor |
|---|---|
| Circos Visualisation | Helena Rasche |
| Genomic Data Visualization with JBrowse | |
| Visualization of RNA-Seq results with Volcano Plot in Galaxy and R | Maria Doyle |
Galaxy for Non-Genomics
Galaxy is widely used for analysis of genomics data, but is not limited to any scientific domain. This module covers some non-genomics topics such as Climate research and cheminformatics.
| Topic | Presentor |
|---|---|
| Climate: Functionally Assembled Terrestrial Ecosystem Simulator (FATES) | |
| Cheminformatics: High Throughput Molecular Dynamics and Analysis | Simon Bray, Chris Barnett |
| Cheminformatics: Protein-ligand docking | |
| Cheminformatics: Running molecular dynamics simulations using GROMACS | |
| Ecology: Compute and analyze biodiversity metrics with PAMPA toolsuite |
All other Tutorials
Please also feel free to explore all the other tutorials on the GTN website, and do any that sound interesting to you. Our instructors will do their best to support you in any of these tutorials.
All done?
Please feel free to hang around in Slack and talk to us and the rest of the Galaxy community! Thanks for joining!!
| Topic | Presentor |
|---|---|
| Wrap up & Socialize | |
| Useful links: Stay involved in the Galaxy Community! | The Global GTN Community |
| Feedback Survey & Certificates |
After the Course
All these materials will remain online, so you can continue working on them for as long as you want. The only difference will be that you should ask your questions on the GTN Gitter channel, instead of Slack.
Admin Track: Overview table
Introduction
Start here. This will cover all the logistics and practical information for this training week.
| Topic | Presentor |
|---|---|
| Welcome to GCC2021 & Course Information | Helena Rasche, Saskia Hiltemann |
| Admin Training Logistics | Helena Rasche |
| About Certificates | |
| Meet & Join the Galaxy Community! | The Global GTN Community |
| Webinar Series: Galaxy Resources For.. |
Introduction to Galaxy
Start here if you are new to Galaxy. These videos will introduce you to the Galaxy platform, and walk you through your first analyses
| Topic | Presentor |
|---|---|
| A Very Short Introduction to Galaxy | Anton Nekrutenko |
| Galaxy Intro for Everyone (non-genomics) | Anne Fouilloux |
Day 1
| Topic | Presentor |
|---|---|
| Daily Icebreaker Question | |
| Ansible | |
| Installing Galaxy with Ansible | Helena Rasche |
Day 2
| Topic | Presentor |
|---|---|
| Daily Icebreaker Question | |
| Running jobs in Singularity | Simon Gladman |
| Ephemeris | Catherine Bromhead |
| Users, Groups, and Quotas | These slides are narrated by AWS Polly. |
| Reference Data | These slides are narrated by AWS Polly. |
| Data Libraries | Saskia Hiltemann |
Day 3
| Topic | Presentor |
|---|---|
| Daily Icebreaker Question | |
| Galaxy Cluster Computing | Helena Rasche |
| Mapping Jobs to Destinations | Helena Rasche |
Day 4
| Topic | Presentor |
|---|---|
| Daily Icebreaker Question | |
| Pulsar | Simon Gladman |
| DB queries, command line & scripts | These slides are narrated by AWS Polly. |
| Monitoring: Telegraf, InfluxDB, Grafana | |
| Maintenance, Backup and Restore | These slides are narrated by AWS Polly. |
| Upgrading Galaxy | Simon Gladman |
Day 5 - Choose Your OWN Adventure
| Topic | Presentor |
|---|---|
| Daily Icebreaker Question | |
| Meet & Join the Galaxy Community! | The Global GTN Community |
| Monitoring With Reports | |
| Training Infrastructure as a Service (TIaaS) | Helena Rasche |
| Interactive Tools | Anthony Bretaudeau |
| Jenkins & Automation | |
| Advanced Customisation | |
| When things go wrong: Galaxy Server Troubleshooting | |
| Python 2 to Python 3! | |
| Tool Development | |
| Dataset Collections | |
| Developing your own Training | |
| Storage management | Gianmauro Cuccuru |
Developer Track: Overview table
Welcome & Practical Information
Start here. This will cover all the logistics and practical information for this training week.
| Topic | Presentor |
|---|---|
| Welcome & Course Information | Helena Rasche, Saskia Hiltemann |
| Get set up for the course | |
| About Certificates | |
| Meet & Join the Galaxy Community! | The Global GTN Community |
| Webinar Series: Galaxy Resources For.. | |
| Daily Icebreaker Questions |
Galaxy Tool Development
This module covers everything you need to start developing your own Galaxy Tools. This module will likely take you several days to complete.
| Topic | Presentor |
|---|---|
| Galaxy Tools: From Conda Through Deployment | Alex Ostrovsky |
| ToolFactory: Generating Tools From Simple Scripts |
Galaxy Training Material Development
Here you can learn about contributing to the GTN’s development, adding new tutorials or slides, and correcting any typos you might spot in the training materials this week.
| Topic | Presentor |
|---|---|
| The GTN & Training Material Development |
Galaxy Core Development
Ever wanted to contribute features to Galaxy? Here you’ll learn a bit about how Galaxy works and how to debug issues in features in Galaxy.
| Topic | Presentor |
|---|---|
| Debugging Galaxy |
Galaxy Architecture Lectures
Here you’ll learn about the internals of Galaxy in a lecture from John Chilton
| Topic | Presentor |
|---|---|
| Galaxy Architecture | John Chilton |
Self-Study Galaxy Development
These tutorials don’t have video walkthroughs, but will all be supported by their authors during the week. So please work through these tutorials by yourself, and ask LOTS of questions in Slack!
| Topic | Presentor |
|---|---|
| BioBlend: Scripting Galaxy using the API | |
| Contributing to BioBlend Development | |
| Contributing extensions to Galaxy core |
Acknowledgements
This Global Galaxy course is only possible thanks to a Global network of instructors and institutes.
Presenters & Instructors & Facilitators & Community Caption Contributors
Institutions