Development News Brief
On this page
Get Galaxy
-
new:
% hg clone http://www.bx.psu.edu/hg/galaxy galaxy-dist -
upgrade:
% hg pull -u -r 338ead4737ba
News
* Keep current on all things Galaxy at our [News](/news/) and [Events](/events/) wikis. * Galaxy is still [hiring](/galaxy-is-hiring/)!What’s New
Galaxy Main
- Data and Job Quotas announced earlier today by email, Galaxy wiki, Twitter, and News Items/RSS feed.
- Email message from the team to galaxy-user and galaxy-dev mailing lists.
- Main wiki with limits, FAQ, and related links.
- Managing Datasets wiki contains detailed help to organize, clone, copy, save, and delete unneeded data.
- Disk Quotas wiki has administrative information on how Galaxy’s disk space accounting works, how to keep it up to date, and how to enable and manage quotas
- Twitter #galaxyproject.
- Galaxy News Items and RSS feed.
User Interface (UI)
- Add the ability for a user to log out of all user sessions. Useful when sharing computers, using public computers, and the like.
- Located under top menu bar: User -> Preferences ->Logout of all user sessions.
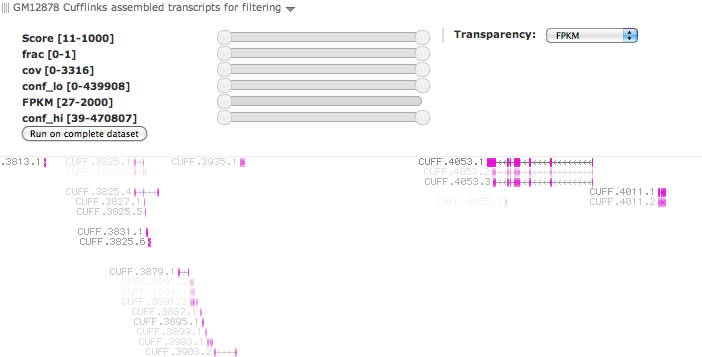
Trackster (Galaxy Track Browser)
- The October 23-24, BioVis 2011, Providence, Rhode Island, United States, included a talk covering the visualization tool Trackster aka Galaxy Track Browser (GTB). Abstract is here.
- Overview provides chromosome-wide view of a single track independent of primary, interactive view.
- Make it possible to set features’ transparency based on attribute values.
- Introduce track groups, which can be used to cluster and move tracks together. Running tools places new tracks in groups.
- Enable tools that produce BED output to be used in visual analytics framework and enable GOPS cluster tool to work in Trackster.
- Use Fasta datasets to specify custom build visualizations.
Python and OS Support
- Python 2.7 is moved from beta to fully supported, any issues with Python 2.7 should be reported and will be addressed.
- Python 2.4 support is officially deprecated and will only be maintained through this stable release and the release following it.
- Eggs are now provided for versions of Python 2.7 for Mac OS X downloaded from python.org with platform string ‘py2.7-macosx-10.6-intel-ucs2’ (10.6 is the minimum version supported, the platform string is the same on 10.7).
Galaxy Tool Shed
* [Search repositories for valid tools by any combination of id, name or version](/toolshed/searching-the-toolshed/#search-repositories-for-valid-tools-by-any-combination-of-id-name-or-version). * [Automatic installation of Galaxy tool shed repository tools into a local Galaxy instance](/admin/tools/add-tool-from-toolshed-tutorial/). * [Getting updates for tool shed repositories installed in a local Galaxy instance](/updating-installed-repositories/#getting-updates-for-tool-shed-repositories-installed-in-a-local-galaxy-instance). * Fixes for displaying file names that include spaces when browsing repository contents.Updated & Improved
Tools
- Add GTF-guide option to the Cufflinks wrapper.
- Add wrapper for Trinity tool, the de novo RNA-seq transcript assembler. Available for local or cloud instances only.
- Add wrapper for SICER tool, a Statistical approach for the identification of ChIP-Enriched Regions.
- Add wrapper for SAMtools rmdup tool, which removes potential PCR duplicates in BAM files.
- Add the tool Text Manipulation -> Secure Hash / Message Digest:
- Update the Text Manipulation -> Select random lines from a file tool:
- does two passes through input file: first pass determines line offsets/count, offsets are randomly removed, second pass writes out lines by seeking to remaining offsets.
- the random seed can now be optionally specified.
- New assertion-based tests for tool test outputs from John Chilton. For usage, see the example in the Tool Config Syntax documentation.
Quotas and Data Management
- Various performance and usability enhancements, as well as a fully implemented API for the quota system.
- Documentation at Disk Quotas and Managing Datasets.
- Purged histories were counting in users’ disk usage calculations, this has been fixed, so you should rerun
galaxy-dist/scripts/set_user_disk_usage.pyto recalculate usages. - The time to load the list of saved histories should be greatly improved due to a more efficient query to calculate the sizes of histories in the list
- It’s now possible to purge all datasets in a history which are already deleted, as well as purge entire histories that are already deleted.
- Purged histories are now visible in the saved history list, and any history can be viewed before being switched to.
Source
- Tools that include parameters that are ‘dynamic select lists’ that refer to local index files (
.locfiles) or entries in thetool_data_table_conf.xmlfile will now all successfully load even if these dependencies are missing. However, an error message will be displayed in the tool form for these tools conveying information about the missing dependencies. - Framework for refactoring the UI and actions (implemented by the Quota interfaces, new interfaces should also use this method):
- UI components now live in subclasses of
BaseUIController and BaseAPIController, themselves subclasses of BaseController. - Actions, i.e. the work performed by choices made in the Web UI or API now live in galaxy.actions, so they can be called cleanly from either interface
- Parameters passed in the Web UI or API should be parsed in galaxy.web.params. If standardized across all forms and API methods of a particular model class, these parsers can be generalized quite easily
- UI components now live in subclasses of
- Refactor validation of user account data—email, username, password—into own module and use module for validating user data throughout Galaxy.
- Improve GTF sniffing by doing proper parsing of attributes.
API
- Fully implemented API for managing disk quotas.
- Disk usage now included in user API output.
- API for assigning roles to a data library from John Chilton.
- Encoded IDs of library contents in the Library API are no longer the encoded version of file.<library_dataset.id> and folder.<library_folder.id>. Instead, the real ID is encoded and if the library content is a folder, the encoded ID is prepended with an ‘F’.
@web.require_admindecorator can now be used on API controller methods.- For
galaxy.model.<model_class>.get_api_value(),api_*_visible_keyscan now contain names that are mapped attributes to other model classes (thus allowing forget_api_value()recursion to other classes). - For model classes that can be //marked// deleted but are not actually deleted, a new route mapper can be used which makes deleted objects accessible via their own collection (for details see changeset/ca6db8f67477).
- New generic scripts
create.py,delete.pyandupdate.pyingalaxy-dist/scripts/api/to performPOST,PUTandDELETEoperations in the API (useful for development and debugging). - Exceptions in the API are now logged instead of passed through to the client.
- API makes much better use of HTTP response codes now.
Scripts
- “Galaxy ls” from
Simon McGowanadded to thegalaxy-dist/contrib/directory allows users at sites where Galaxy users match system users to get a list of filesystem paths to their Galaxy data (see the in-file Perl documentation for usage)
Bug Fixes
- Tasked jobs will now inherit the parent job’s user information correctly.
- Remove unspecified build validators from Cuffdiff wrapperfixes https://bitbucket.org/galaxy/galaxy-central/issue/631.
- Remove unspecified build validator from extract_genomic_dna tool (allows custom reference genomes).
- Fix bug in setting Cufflinks’ reference annotation.
- Fix bug in cloning a visualization.
- Fix problems with computing indices for GFF/GTF datasets.
- No longer count purged histories in users’ disk usage
- Fix Python 2.4 incompatibility broken in the last stable release
- The DRMAA job runner incorrectly included ’#-S /bin/bash’ in the job template. This is SGE-specific and should be placed in ~/.sge_request if needed at your site.
About Galaxy
The GalaxyTeam is a part of BX at Penn State, and the Biology and Mathematics and Computer Science departments at Emory University.
Galaxy is supported in part by NSF, NHGRI, the Huck Institutes of the Life Sciences, and The Institute for CyberScience at Penn State, and Emory University.