Development News Brief
On this page
All News Briefs
Distribution Summary 2014_06_02
Get Galaxy
 |
getgalaxy.org | |
| galaxy-dist.readthedocs.org | ||
| bitbucket.org/galaxy/galaxy-dist | ||
| new: | $ hg clone https://bitbucket.org/galaxy/galaxy-dist#stable |
|
| upgrade: | $ hg pull $ hg update latest_2014.06.02 |
|
## Upgrades
- Review instructions for core build: Getting the Stable Distribution
- Review reset for Tool Shed repos: Resetting Metadata For Installed Repositories
## Upgrade Notice: latest_ tags
With this release comes the inclusion of new Mercurial tags designed to make the life of a Galaxy server administrator easier. The recommended tag to use for this release is now latest_2014.06.02. This tag will always point to the latest stable version of the 2014.06.02 release (we publish occasional bug fixes to stable releases). If you would like to ensure you always run a static release version with no additional changes since the release, you may use the traditional release_2014.06.02 tag (e.g. hg update release_2014.06.02). Both release_ and latest_ tags will be available for all releases going forward, and a latest_2014.04.14 tag was created for the latest stable version of the previous release.
## Upgrade Notice: "ucsc_build_sites.txt"
If you have made local modifications to the tool-data/shared/ucsc/ucsc_build_sites.txt file, you will need to exercise some caution when upgrading to this release (the easiest solution is to make a backup copy of the file prior to upgrading). See Pull Request #340 for more detailed instructions. There is also a related trello card.
## Dataset Collections
Dataset Collections (Technology Preview): Galaxy’s data model, API, tool and workflow subsystems have been updated to incorporate the concept of dataset collections. Outlined in these 12 trello cards, dataset collections allow performing complex analyses over lists of dataset, paired datasets, lists of paired datasets, etc…
We are calling these a technology preview in this release because there is currently no rich user interface for creating or viewing dataset collections and there are a number of known issues. Check out this presentation, scheduled to given at this year’s Galaxy Community Conference, for the most up to date information on dataset collections and example usages.
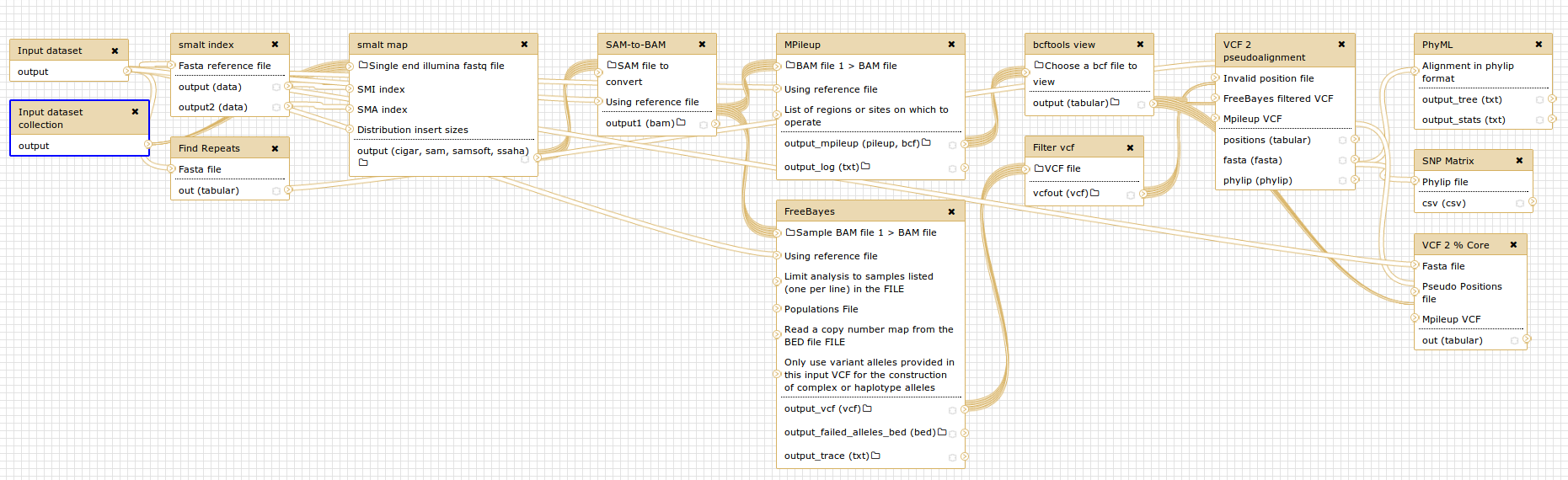
The workflow image above (created by Philip Mabon) demonstrates the usage of dataset collections in the context of the [Core Phylogenomics SNP](https://github.com/apetkau/core-phylogenomics) pipeline created by Aaron Petkau, Gary Van Domselaar, Philip Mabon, and Lee Katz.
## Pull Requests Merged
You will now find merged pull requested incorporated directly into the News Brief along side other changes. Thanks to our Galaxy community contributors!
## Galaxy Biostar
If you haven’t claimed your account yet at Galaxy Biostar, now is a good time. This is the last reminder that the galaxy-user@bx.psu.edu mailing list will be retired Friday, June 6th. https://trello.com/c/0nQ5tHG7
## Data
- Data Manger:
- Genome Builds / dbkeys: Make adding builds accessible by Data Manager. tools https://trello.com/c/5d077rYP
- Reference Data:
- Replaced: hg19 100waymultiz conservation alignments sourced from UCSC utilized by “Fetch Alignments (MAF)” tools and available via rsync. https://trello.com/c/hlNAx4wq https://trello.com/c/5d077rYP
- Updated: NCBI taxonomy data utilized by metagenomics analysis tools and available via rsync. https://trello.com/c/5d077rYP
- Updated: NCBI database indexes utilized by Megablast and available via rsync. https://trello.com/c/jH2UBpaJ
- Correction: Error in alignseq.loc entry for “Arabidopsis_thaliana_TAIR10”. https://trello.com/c/oEgtg6yc
- Documentation: Wikis for data prep reorganized and updated. Start here…
## Tools
- Execution: Multi-dataset execution (a la workflows). https://trello.com/c/14OxiyMv
- Default value for
data_columnnot working in tool XML. Peter Cock PR#389. https://trello.com/c/6ugfcLgI - Provide mechanism to cleanly inject arbitrary environment variables from destinations (static and dynamic) into job script. PR#378. https://trello.com/c/t1FH1Q5P
- Enhancements for configuring tools that produce a variable number of outputs. PR#356. https://trello.com/c/3SNHcQlU
- Pull in a complete tool input parameters spec in JSON (e.g. recursion into Group parameters: Conditional and Repeat). https://trello.com/c/ydPxchL4
- Toolshed Tools: Display link to toolshed repository on form. https://trello.com/c/QRAP0eFj
- Changed
selected_filetorepo_file. ResolveNameError: global name ‘selected_file’ is not defined. Trevor Wennblom, PR#382. https://trello.com/c[/FhofbMxp](/archive/dev-news-briefs/2014-06-02/FhofbMxp/)
## Visualizations
- Trackster:
- Implement location router in shared visualizations. https://trello.com/c[/C7Ypv](/archive/dev-news-briefs/2014-06-02/C7Ypv/)TAi
- Put converted datasets into history as hidden dataset. https://trello.com/c[/G5ZzUb3](/archive/dev-news-briefs/2014-06-02/G5ZzUb3/)X
- External Display Applications:
- Updates to IGB display of XML files. https://trello.com/c/e23KKCKb https://trello.com/c/qJyALlN7
- Added IGB display support for bed graph files. hillrunner2008, PR#361. https://trello.com/c/pJeuhEP3
GenomeSpace(OpenID) Token Expiration. https://trello.com/c/riMxlQ6k
## Workflows
- Workflow’s with tool steps utilizing multiple input data params no longer break when edited (reported by Peter Cock). https://trello.com/c/9knsbvmx
- Corrected problematic backtrace when attempting to run workflow after upgrade. https://trello.com/c/zXkliR75
- Make workflow URLs pretty and establish conventions. https://trello.com/c/nvR6hktq
- Workflow editor now preserves conditionals (avoids a specific invalid state that previously broke connections). https://trello.com/c/4yWqzlRi
## UI
- Dataset provenance functionality improved for all user types. https://trello.com/c/ms6rtKmQ
- Improve public
usernamehandling at registration (including via API). https://trello.com/c/UAfb8SVX - Improve
History options -> Purge deleted datasetsto update user disk usage. https://trello.com/c/bvOL9IBb - Simplify localization of output and ensure acceptance of UTF-8 in history, dataset API, and client code. https://trello.com/c[/SxVszv0r](/archive/dev-news-briefs/2014-06-02/SxVszv0r/)
- Create a proper JS object to contain components (user, configuration, logger). https://trello.com/c/JXPol7vs
- Extending the UI library: new elements. https://trello.com/c/xYiHvOfl
## API
- API’s authenticate/baseauth creation of API key. https://trello.com/c/zpK846FE
- Allow fetching input/output dataset association information through jobs API. https://trello.com/c/1ly1QN3a
- Include job id in dataset provenance API. https://trello.com/c/SH8mWi8J
- Expose use_remote_user via configuration API. https://trello.com/c/AGKePuHZ
- Update code checks and documentation after commit 17a2dc7. Nicola Soranzo, PR#383. https://trello.com/c[/Ms6E0enr](/archive/dev-news-briefs/2014-06-02/Ms6E0enr/)
- Fix for “‘HistoryTagsController’ object has no attribute ‘security_check’”. Nicola Soranzo, PR#394. https://trello.com/c/7iIv01zm
## Core
- Admin:
- Messaging and Task Queue update. https://trello.com/c[/B0pV80d0](/archive/dev-news-briefs/2014-06-02/B0pV80d0/)
- Client side logging for metrics and API entry point for fluentd. https://trello.com/c/7yfdzZj2
- Improve
View data tables registryfor developers. https://trello.com/c/gmClxfV8 - Update
phantomjs 1.9andcasperjs 1.1plus tests and modules intest/casperjs. https://trello.com/c/VjDIpgt3 - Page controls now loaded with
history/list_published.mako. https://trello.com/c/nR2X7VJW - Added rolling restart script to kill servers sequentially. Helena Rasche, PR#393. https://trello.com/c/jdldxhlk
- Config:
- Admin action to reload “.loc” files without restarting Galaxy. https://trello.com/c/c7nUco3u
- Allow for an arbitrary number of
tool_data_table_config_pathfiles. https://trello.com/c/rm2p8PpZ - Better reflect
allow_user_dataset_purgeconfiguration in history panel. https://trello.com/c/RCPZ9zMF - Create
run_tests.shto replacerun_functional_tests.sh. https://trello.com/c/46KduuJs - Remove misc files that slipped past
hgignore. Trevor Wennblom, PR#364. https://trello.com/c/NlbwJMjA
- Jobs:
- Documentation Updates: https://trello.com/c/jDLtI38Z https://trello.com/c/CRjWSodv
- https://wiki.galaxyproject.org/Admin/Config/JobMetrics
- https://wiki.galaxyproject.org/Admin/Config/Jobs#Job_Destinations
- job_conf.xml.sample_advanced
- Plugin framework and plugins for job metrics. https://trello.com/c[/MexBahA3](/archive/dev-news-briefs/2014-06-02/MexBahA3/)
- Capture and report time taken to run each job. https://trello.com/c/tRxCl9xv
- Local Job Runner Enhancements. PR#384. https://trello.com/c/PDwEzOj1
- Improve
run_functional_tests.shscript. https://trello.com/c/14Mt7Nap - Fix GALAXY_SLOTS calculation for LSF. Ilya Sytchev. https://trello.com/c/WZ7ydlnW
- Add option test system environment option GALAXY_TEST_VERBOSE_ERRORS to report on dataset errors. https://trello.com/c/neyVEnxZ
- Update the job exit codes. Jian-Long Huang, PR#365. https://trello.com/c/BXldAhSn
- CLI Job Runner Enhancements. PR#385. https://trello.com/c[/KclOs](/archive/dev-news-briefs/2014-06-02/KclOs/)ZJi
- Change:
job_directory -> jobs_directory. Trevor Wennblom, PR#360. https://trello.com/c/DXBL1XTt - LWR Message Queue Fixes and Improvements. https://trello.com/c/wMvq2qNZ
## Security
- Change to avoid potential privacy exploit during login. https://trello.com/c[/XlXwM866](/archive/dev-news-briefs/2014-06-02/XlXwM866/)
- Remove use of unencoded ids from history_common.mako, related controllers. https://trello.com/c/psDNnyRL
Fixes
- Fixed issue with ‘bug report’ submission using the Google Chrome web browser. https://trello.com/c/SX1WSchz
- Fixed library permissions API controller. https://trello.com/c/80EmYp19
- Fixed ‘multiple input data parameters’ in workflows. https://trello.com/c/0neRZzj9
- Fixed library permissions API controller. https://trello.com/c/80EmYp19
- Replaced loading indicator for Tabular View. https://trello.com/c/rP8xrqvH
- Fixed problem when uploading to a Library does not correctly generate external set_metadata command (auto-detect metadata also broken). https://trello.com/c/25XLSk0r https://trello.com/c/95t7B3EV
- Fixed the column alignment in the tabular format data. Jian-Long Huang, PR#366. https://trello.com/c/LXnPuIRR
- Fixed tag editing in Pages. https://trello.com/c/Yg7YGm6p
- Fixed ‘multiple input parameters’ causing repeated datasets in UI names. https://trello.com/c/LCEKxImR
- Fixed
ToolShedinstallation using virtualenv that failed to automatically configure paths. https://trello.com/c/VFHrmuOS - Fixed
<conversion>tags within aDataToolParameterthat no longer worked. https://trello.com/c/9diH0RFl - Fixed grid templates that were inserting whole documents inside other documents. https://trello.com/c/iYloJMUF
- Fixed problem with primary dataset collection on cloned histories. https://trello.com/c/17McdFlx
- Fixed issue with Biostar “forgotten password” reset. https://trello.com/c/AX6zraeC
- Misc improvements: Remove unused methods, cleanup imports, fix old wiki links. Nicola Soranzo, PR#362. https://trello.com/c/5dE7l7YC
## Tool Shed
Repositories
- Add
assert file existsanddirectory existsactions totool_dependencies.xml. https://trello.com/c/xBUhFvj0 - Making it easier to package tools using a local toolshed. https://trello.com/c/xtA7DsTz
- Display mercurial ctx revision number instead of hash string. https://trello.com/c/ldVAUyYL
- Reserved words
$REPOSITORY_INSTALL_DIRand$INSTALL_DIRcan now be used in certain tool actions such as ‘move_file’ and ‘move_directory’. https://trello.com/c/XK5SqE1i
API
- Improve API error message when sent an incorrect user name. https://trello.com/c/bxZiluH3
- API installation of “Repository suite definition” no longer fails when there are multiple, different, owners. https://trello.com/c[/Bxe2E5Pp](/archive/dev-news-briefs/2014-06-02/Bxe2E5Pp/)
- Enhance the API to provide the current set of categories. https://trello.com/c/xaUy82rl
Test Framework
- Better handling of tool test results for tools missing tool test components on test tool shed. https://trello.com/c/rArkn49z
- Install and test framework Stage 2 now runs. https://trello.com/c/YElEsCWl
Tool Shed Fixes
- Fixed missing searchbar in Test Tool Shed via
FireFoxbrowser. https://trello.com/c/JgdgdgOH
Project Updates
## About
The Galaxy Team is a part of the Center for Comparative Genomics and Bioinformatics at Penn State, and the Department of Biology at Johns Hopkinis University.
Galaxy is supported in part by NSF, NHGRI, the Huck Institutes of the Life Sciences, and The Institute for CyberScience at Penn State, and Johns Hopkins University.
The public Main instance of Galaxy at http://usegalaxy.org utilizes infrastructure generously provided by the iPlant Collaborative at the Texas Advanced Computing Center, with support from the National Science Foundation.
Follow us on Twitter @galaxyproject or read our tweets at Galaxy on Twitter
Have a suggestion? Please see Using the Galaxy Issue Board, and be sure to up vote the issues important to YOU.

