New tutorial on selection of candidate biomarkers
Featuring the ProteoRE server
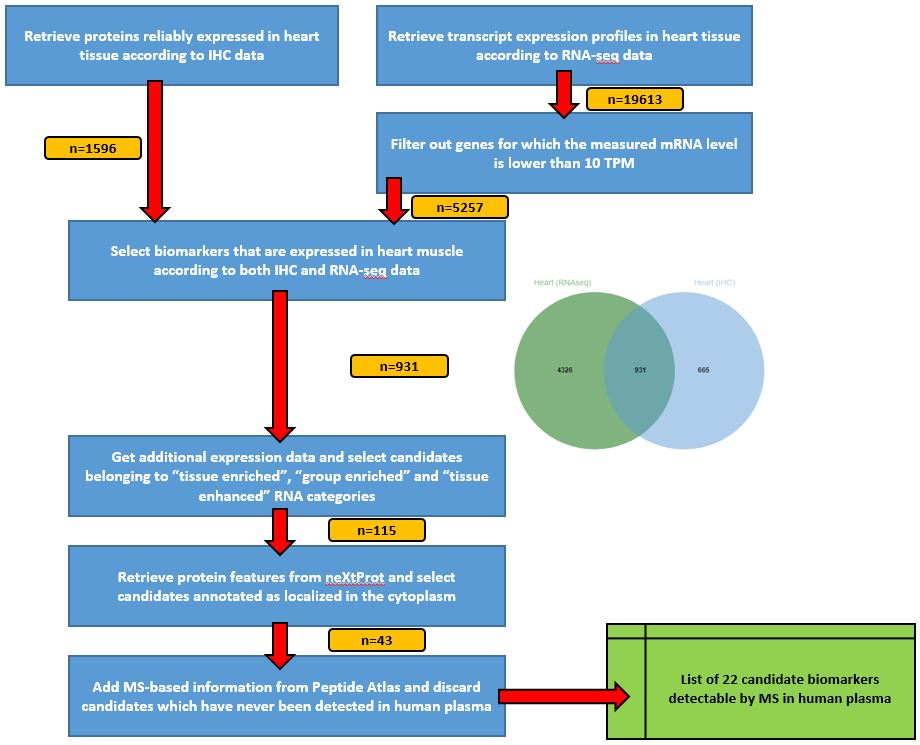
The ProteoRE team is pleased to announce the publication of a new tutorial in the Galaxy Training Network directory. As a new training material for proteomics workflows in Galaxy, this tutorial is dedicated to the selection of candidate biomarkers; it illustrates how to use ProteoRE’s tools to design a strategy in a stepwise manner for reproducible biomarker selection relating to disease pathophysiology. In line with GTN’s recommendations and thanks to their assistance, particular attention has been paid to make this training material suitable for non-expert biologists and for self- study, explaining everything on the biological level (i.e. what is the data we have? what are the relevant biological concepts to understand? why are we doing these steps? what do the results tell us? etc).
ProteoRE (Proteomics Research Environment) is a biologist-oriented web platform for biomedical research. Built upon the Galaxy framework, ProteoRE provides access to tools, workflows, and data for the functional analysis and the exploration of proteomics and transcriptomics data.
Questions? Contact the ProteoRE Team contact@proteore.org