COVID-19
| Public server: | Devoted to research on coronavirus disease 2019 (COVID-19) and SARS-CoV-2 |
|---|---|
| Scope: | Domain specific |
| Summary: | A companion to our study describing the analysis of early COVID-19 data |
comments
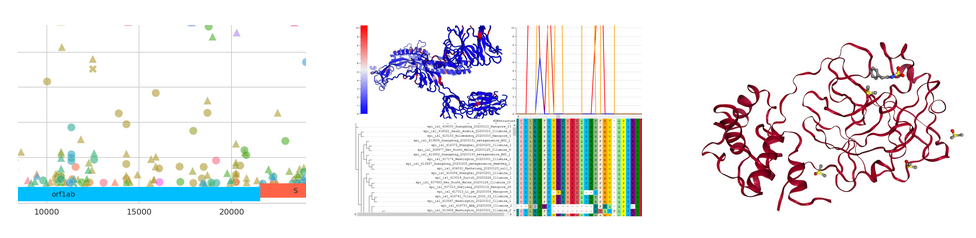
This subdomain servers as a companion to our study describing the analysis of early COVID-19 data, the goal of which is to underscore the importance of access to raw data and demonstrate that existing community efforts in curation and deployment of biomedical software can reliably support rapid reproducible research during global crises.
- This subdomain contains the exact versions of all software used.
- Our analysis was divided into six parts. Each part has a dedicated page that provides links to input datasets, intermediate and final results, workflows, and Galaxy histories that list all details for each analysis. These workflows can be re-run on this subdomain.
- The public server is hosted by the UseGalaxy.eu team.
user support
quotas
Storage and computational quotas.
citations
Baker, D., Beek, M. van den, Blankenberg, D., Bouvier, D., Chilton, J., Coraor, N., Coppens, F., Eguinoa, I., Gladman, S., Grüning, B., Keener, N., Larivière, D., Lonie, A., Pond, S. K., Maier, W., Nekrutenko, A., Taylor, J., & Weaver, S. (2020). No more business as usual: Agile and effective responses to emerging pathogen threats require open data and open analytics. PLOS Pathogens, 16(8), e1008643. DOI: 10.1371/journal.ppat.1008643
sponsors
The Freiburg Galaxy Team but also collectively by groups and individuals from across Europe