Development News Brief
On this page
Get Galaxy
Please note new upgrade syntax
 |
getgalaxy.org | |
| galaxy-dist.readthedocs.org | ||
| bitbucket.org/galaxy/galaxy-dist | ||
| new: | $ hg clone https://bitbucket.org/galaxy/galaxy-dist#stable |
|
| upgrade: | $ hg pull $ hg update release_2013.02.08 |
|
## Improvements to Distribution Release Process
THIS IS IMPORTANT, PLEASE READ BEFORE UPGRADING
We have made some improvements to our release process, and because of this, you must include the new distribution release tag in your hg update command to upgrade to this distribution release:
% hg pull
% hg update release_2013.02.08If you do not include the release_2013.02.08 tag, your repository will update to the default branch, which includes unstable code. After updating, you can verify that you are on the stable branch with:
% hg branch
stableOnce you’re on the stable branch, hg pull -u without a specific revision or branch will cause you to remain on the stable branch.
The new release process works as such:
- The Galaxy source repository now contains two branches:
stableis committed to regularly and contains important bugfixes.defaultis committed to regularly and contains new features as well as fixes merged fromstable.
- Every two months,
stablewill be updated with new features from default and a specific distribution release version will be tagged. - The two weeks prior to the release will be focused on fixing bugs found in the targeted upcoming release. New features will not be considered for inclusion in the upcoming release during this time.
This process will not only allow us to deploy important bug fixes on to the stable branch rapidly and in between releases, it will improve the quality of our tagged releases. The documentation at our wiki reflects these changes: getgalaxy.org.
## Tool Shed
New Tool Shed Features
Complex repository dependencies: tool dependency definitions that contain repository dependency definitions
Complex repository dependencies allow for many tool shed repository installations into a single Galaxy instance where any number of the contained tools across installed repositories can all reference a single installation of a specific 3rd party tool dependency (e.g. bwa version 0.5.9).
Describing how this works is tricky, so let’s see if we can state it in a way that can be understood: tool dependency definitions that contain repository dependency definitions define a relationship to a tool dependency that will be installed using the instructions in the required repository’s tool_dependency.xml file, where the required repository is defined by the contained repository dependency definition. Hmmm…okay, let’s try to clarify this a bit… Read more…
The Tool Shed now Provides Sharable URLs for Repositories by Name or Owner
The following new routes have been added to the Tool Shed:
<tool shed url>/view/<repository owner>
<tool shed url>/view/<repository owner>/<repository name>These allow for URLs that can be cited or shared, and viewing a repository will display it’s sharable link. Here’s an example.

Thanks to Peter Cock for the contribution!
Enhanced Galaxy Features for the Tool Shed
- The ability to select multiple tool shed repositories for simultaneous installation has been reintroduced. This feature was eliminated several months ago to allow for the implementation of repository dependencies. Now choosing either of the options “Search for valid tools” or “Search for workflows” from a tool shed’s pop-up menu as shown here…
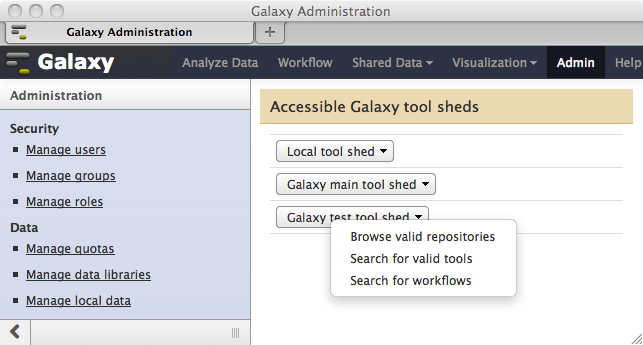
…and entering a search string as shown here…
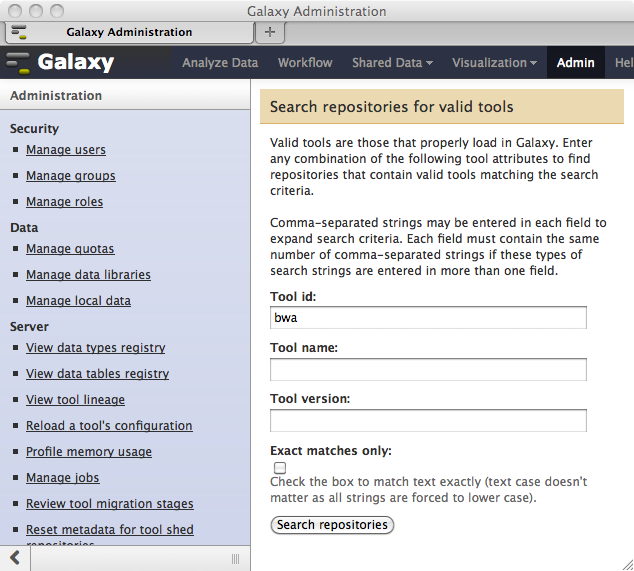
…will allow you to selectively install any of the repositories matching the search criteria.
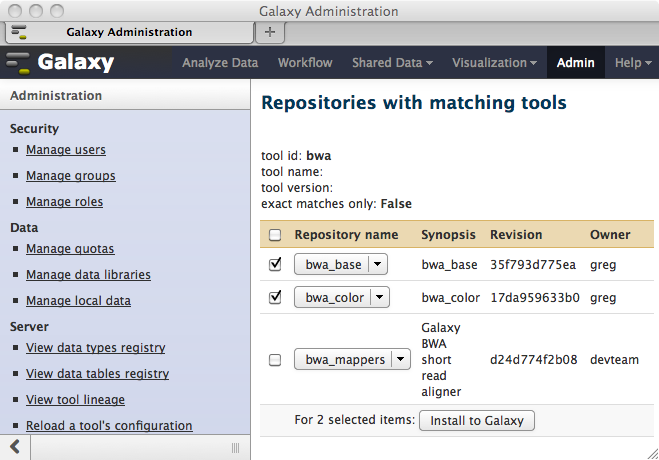
All repositories dependencies and tool dependencies that are defined for each repository selected for installation will be discovered and displayed in preparation for installation into your local Galaxy instance. You can optionally elect to install these dependencies..
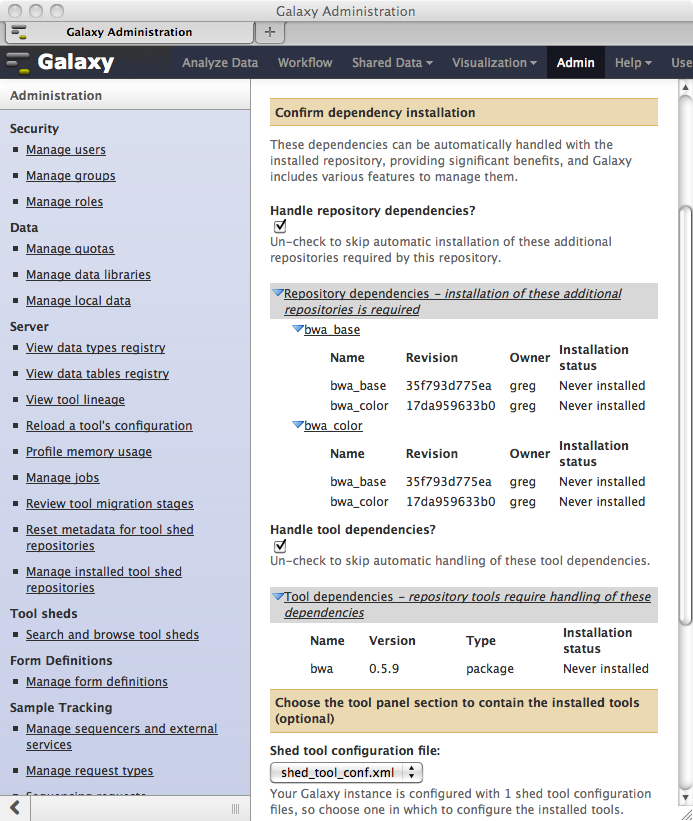
Tool Shed Related Fixes
- Properly handle long URLs when installing and reinstalling tool shed repositories within a Galaxy instance. This issue was reposted by James Johnson, and occurred only when running Galaxy behind an apache proxy, which tends to limit the length of a URL to 8190 characters.
- Add a missing import to the Galaxy
UpdateManger. This issue was reported by Anthonius deBoer, and was corrected inchange set 8649:34404f848979. The GalaxyUpdateMangeris responsible for determining if any updates are available from the tool shed for any repositories installed into the Galaxy instance. The missing import resulted in theUpdateMangernot discovering if updates for installed repositories are available from the tool shed, but the behavior does not result in any instability in the Galaxy instance. The only result of this is that the installed repositories that have updates available are not displayed in yellow highlighted boxes on the “Manage installed tool shed repositories” page within the Galaxy Administration interface. - Invalid
repository dependency definitionsare now properly handled, with error messages displayed within appropriate Galaxy and tool shed features.
Tool Shed Enhancements
- Additional significant enhancements have been made to the tool shed functional tests, so many more tool shed related features are now covered both within the tool shed and with Galaxy.
Miscellaneous Tool Shed Fixes and Enhancements
- Allow repository owners to reset all repository metadata on a specific repository in the tool shed.
- Allow a Galaxy administrator to select a shed-related tool panel configuration file whose tool_path setting will be used to installed repositories into Galaxy for those repositories with which a tool panel selection is not necessary.
- Uncomment the default sqlite database connection string in community_wsgi.ini.sample.
- Automatically create missing required tool shed files from samples when starting the tool shed server.
- Many additional test scripts have been added to the tool shed’s functional test suite.
## Tools
- Enable BedGraph format datasets to be used as input to
wig-to-bigwigtool.
- Renamed tool as:
Wig/Bedgraph-to-bigwig.
- #2 Add parameter to
Filtertool to optionally skip header lines. Add test for new parameter usage as well.
## Data
- Megablast indexes for the divisions
nt,htgs, andwgshave been updated on the public Main Galaxy server dated28jan2013. - NGS Data Set-up wiki updated and simplified. [View here…](http://wiki.galaxyproject.org/Admin/NGS Local Setup)
- Rsync server hosts download of the same
.locfiles used by the Galaxy team (to be used as additional examples, or starter files if you use our rsync reference genome data. Read more…
## CloudLaunch
Update
- Change get_account_info to POST vs GET.
Bug Fixes:
- #2 Adjust event triggers to prevent redundant (though harmless) get_account_info invocations.
- #3 Adjust Cluster List creation to always insert the “New Cluster” option.
## Workflows
- Exporting workflows:
- Remove spurious header.
- Add option to create workflow image.
## Histories
- Enable display icon button for queued, running jobs in the history panel.
- Show API errors for specific datasets as error-ed datasets in the history panel (api/histories/:id/contents?ids=…)
## Admin
- Have
run_functional_tests.shserve static files (images, javascript, etc.) to assist in testing client-side code.
## Framework
- Unified system genome builds, custom builds, build len files, and build two bit files into a single python structure.
- Add function to get genome builds with or without build length information.
- Fix bugs so that the full build set (system + custom) is available via the API and when adding datasets to libraries.
## Source
Pull Requests Merged
- Fixing database v108 downgrade failure, pull request #109. Thanks to Kyle Ellrott.
- Paired-end code mishandles description of FASTQ headers, pull request #8. Thanks to fangly.
- Fix Add/Remove buttons for Repeat groups, pull request #24. Thanks to epaniagu.
- The start of the API based annotation system, pull request #101. Thanks to Kyle Ellrott.
- Add option to bowtie2 tool for writing not aligned reads, pull request #110. Thanks to Nicola Soranzo.
- Adjusted trimming tool for easily trimming from the end, pull request #113. Thanks to Joachim Jacob.
- Adding handler for httpexceptions.HTTPFound, so that the redirect tool will work (rather then causing an error), pull request #92. Thanks to Kyle Ellrott.
- Adding UUIDType to data model and uuid column to dataset table. (This is unused, so far), pull request #105. Thanks to Kyle Ellrott.
- Refactoring Task Splitting Toward Per-Job Definitions (in Addition to Current Per-Tool Definitions), pull request #87. Thanks to John Chilton.
- Fixed error when retrieving workflows via the API, pull request #114. Thanks to Richard Park.
## Security Fixes
Although there are no specific known security fixes in this distribution, all Galaxy instance maintainers are strongly encouraged to run the latest release to take advantage of other improvements and fixes.
## Bug Fixes and related Enhancements
General/UI
- Fix for external display applications with non-public permissions.
- Add missing import in
galaxy.util. Fixes bug in.mkstemp_ln(). Only encountered rarely in the upload tool.
Tools
- #3 Improve error message for tool test parameters that could not be added.
- #4 Add tool id for when clause warnings that are output when tool XML is parsed.
- #5 Use correct index tables for setting dbkey of Tophat2 outputs.
- #6 Fixes for showing job parameters when tools have been updated.
Visualization
- #7 UI fixes for navigating histories/libraries and choosing datasets in Trackster.
- #8 Change ‘clone history/workflow/visualization’ to ‘copy history/workflow/visualization’ for readability
Admin
- #9 A bug has been fixed in the
DRMAArunner that was causing job handler processes to segfault when users deleted history datasets that were associated with jobs that had not yet been dispatched to a cluster. - #10 Adjust the id_tag to use the format “
<jobid>_<taskid>”, instead of “<jobid>:<taskid>”. This resolves issues with particular cluster runners (UGE) being unable to submit.
## Announcements
News, February 2013 Galaxy Update
GCC2013
Training Day Topic Voting Closes 11 February
If you might attend the GCC2013 Training Day then please review the possible topics and then vote for your top 3 choices. Your votes will determine not only the topics that are offered, but also which topics should be offered more than once, assigned to which rooms, and which ones should not be scheduled at the same time. Your vote matters.
Registration, Talk & Poster Submission Opens 22 February
Early Registration, and talk and poster abstract submission all open 22 February. Watch the mailing lists and this wiki for details.
## About Galaxy
The Galaxy Team is a part of BX at Penn State, and the Biology and Mathematics and Computer Science departments at Emory University.
Galaxy is supported in part by NSF, NHGRI, the Huck Institutes of the Life Sciences, and The Institute for CyberScience at Penn State, and Emory University.
Join us at Twitter @galaxyproject or just read our tweets [Galaxy on Twitter](http://wiki.galaxyproject.org/Galaxy on Twitter)
Have a suggestion? Please see Using the Galaxy Issue Board.


