
A fruitful year for the Galaxy Training material
The Galaxy Training Network has developed an infrastructure to deliver interactive training based on Galaxy: one central place (https://training.galaxyproject.org) to aggregate training materials covering many current research topics.
Each topic is supported by different tutorials built to support interactive learning around a "research story" (usually the reproduction of a scientific paper). The tutorials can then easily be used for effective training to support individual users in their self-learning effort and also instructors during workshops.
The tutorials are developed and maintained by the community via a GitHub repository: https://github.com/galaxyproject/training-material.
This project was started in 2016 and was presented for the first time during the 2017 Galaxy Community Conference: Building an open, collaborative, online infrastructure for bioinformatics training. But during the last year, quite a lot has happened with the training material!
Content: New + Restructed Topics and Tutorials
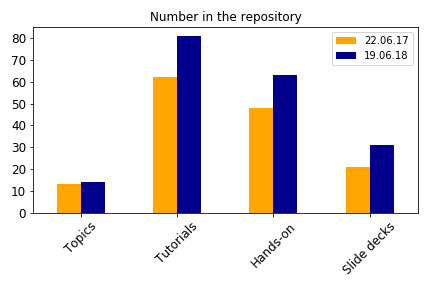
During the past year, two new topics have been added:
While several more new topics are in development (e.g. Visualization, Phylogeny, Metabolomics).
Many existing topics have been expanded with new content:
- Introduction to Galaxy: Extracting Workflows from Histories, Introduction to Genomics and Galaxy and Rule Based Uploader
- Assembly: Unicycler and Making sense of newly assembled genomes
- Epigenetics: Hi-C analysis of Drosophila melanogaster cells using HiCExplorer
- Genome annotation: Genome annotation with Prokka
- Transcriptomics: Differential abundance testing of small RNAs and Visualization of RNA-Seq results with CummeRbund
- Variant Analysis: Mapping and molecular identification of phenotype-causing mutations and Microbial Variant Calling
Some tutorials have been restructured, improved, or updated to use state of the art tools. For example the Reference based RNA-Seq tutorial is now using MultiQC, STAR, FeatureCount and explains how to do a functional enrichment analysis using GOSeq.
The tutorials from the Hub have been migrated to the Training Material repository, placed in the corresponding topics, and formatted to fit the Training material templates.
We also restructured the "Train the Trainers" topic to orient it more to the contribution side. We then developed several new tutorials to help potential contributors learn how to contribute via GitHub or via command-line. Our idea is to help our contributors in the sometimes difficult contribution process. We have also created a Frequently Asked Questions page to answer common questions.
We are still in discussions with the Galaxy Admin instructors to integrate the content of Dagobah with the training material.
More technical support
An effort has also been made to support more the tutorials with their required tools in tools.yaml, workflows, data on Zenodo and referenced in data-library.yaml, and interactive tours:
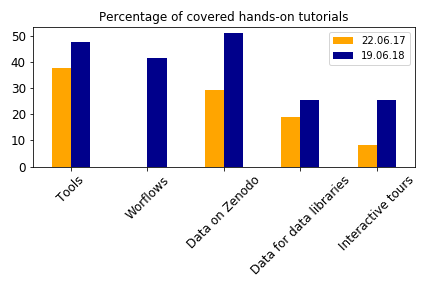
Tours
The development of the interactive tours has been facilitated with the development of the Galaxy Tour Builder: a web-browser extension which allows recording actions in the Galaxy interface in order to create the interactive tours.
Required Tools
The tools.yaml file of each tutorial is used to check which public Galaxy instances could be used to run the tutorial. This information is displayed on the top of the tutorial and in the topic page:
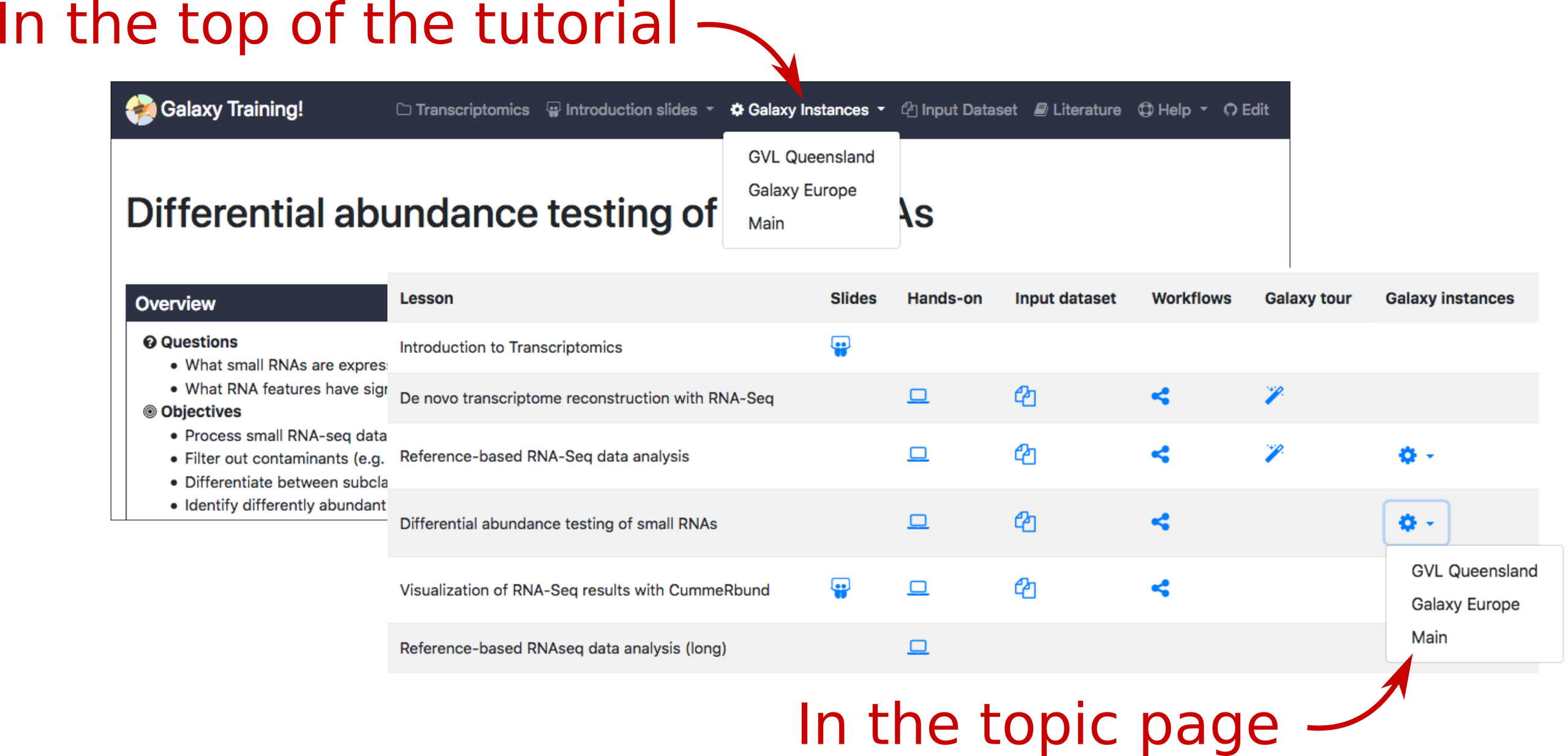
Accompanying the annotation, we also offer for each instance a collection of badges denoting compatibility with training materials (https://training.galaxyproject.org/badges). These badges can then be displayed on the instance homepages.
Training Infrastructure as a Service
The European Galaxy server (http://usegalaxy.eu) is going one step further in the support for training. It offers Training Infrastructure as a Service (TIaaS). An organizer of a training workshop can ask for dedicated computing resources (dedicated queue and nodes on the cluster) for the trainees during the workshop. It is totally transparent for the workshop participants: they uses the UseGalaxy.eu instance as a normal user, but in the backend their jobs are automatically submitted to a dedicated queue. At the end of the workshop, the trainees are placed again on the normal queue:
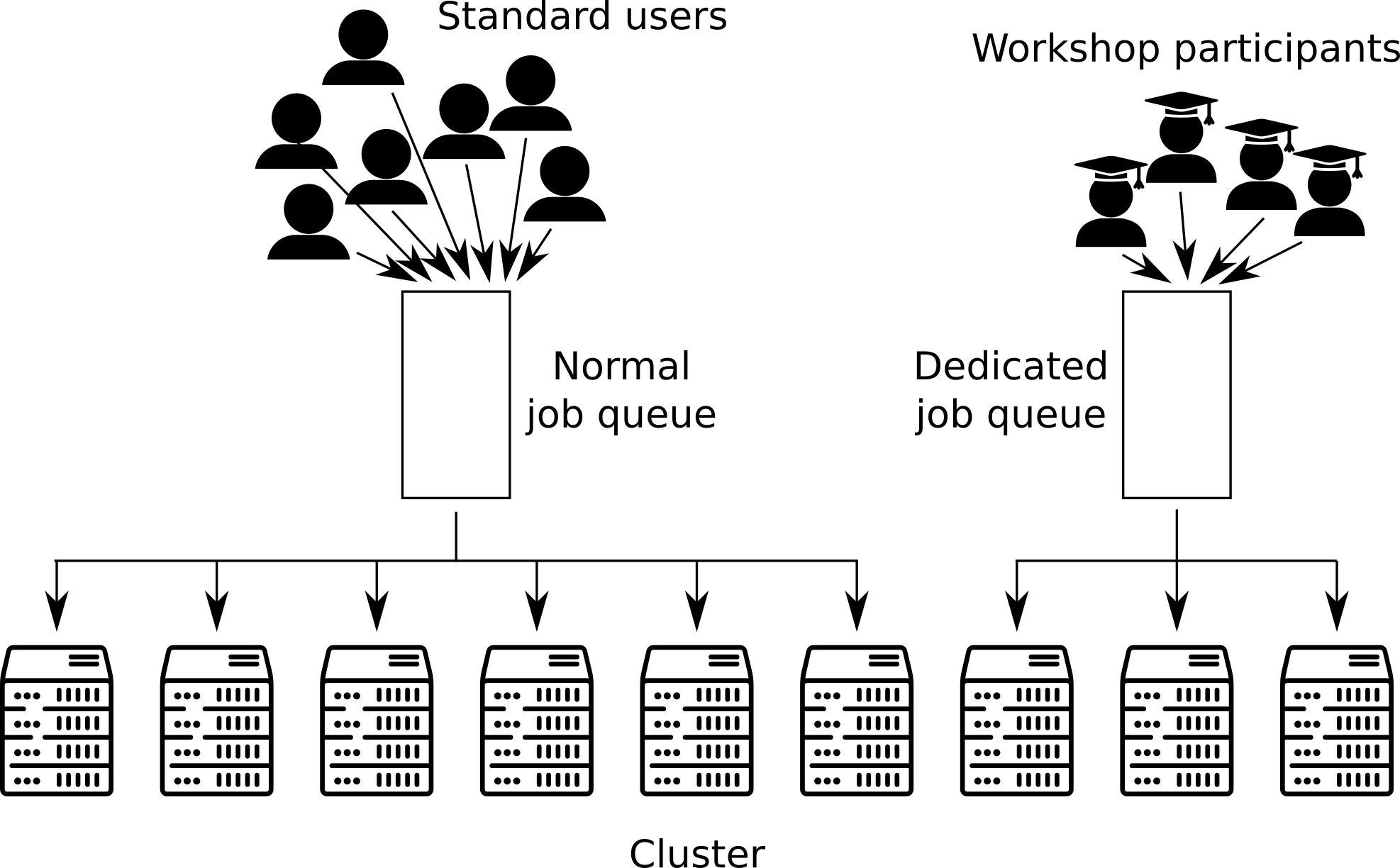
Docker Images
More Docker images are offered to support the training. These Docker images, one per topic, integrate a full Galaxy instance with all the tools, the data, the workflows and the interactive tours to support the tutorials of the topic. They can be easily deployed on a local cluster or cloud.
Improved Website
The website can now be accessed from https://training.galaxyproject.org, which redirects to http://galaxyproject.github.io/training-material. An effort has been made to reshape the website:
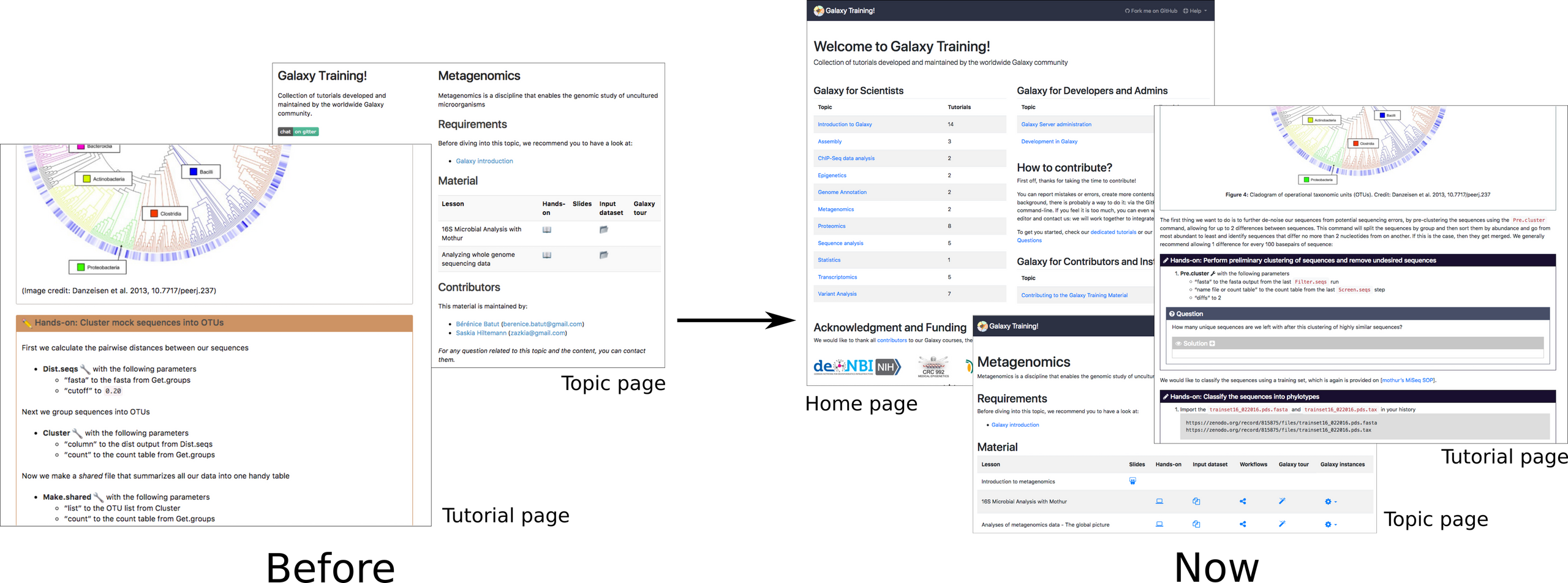
We have also added some expandable boxes for the solutions of the question or more detail boxes. To help in the choice of the tutorials within a topic, we have added the possibility to tag the tutorials directly inside the metadata.yaml. The tags appear on the topic page, right below the title of the tutorial:
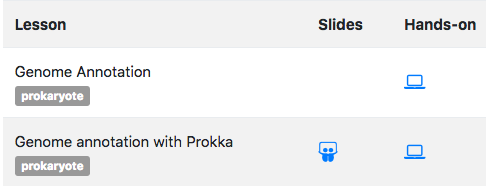
Each training webpage is now annotated with some metadata using the JSON-LD representation of schema.org's CreativeWork vocabulary. This annotation will help the search engines by providing them structured information on our training materials and tutorials, exposing information like title and authorship in a way that they can easily consume and display to their users.
More Support Infrastructure
Over the last year, we have improved the infrastructure. All the material is now correctly referenced on TeSS, the ELIXIR training portal. Any new tutorials or topics will automatically appear there within one day. We also can easily generate PDFs of the tutorials and the slides. Best of all, most of our the training is covered by continuous integration and automatic testing: URLs in the tutorials and slides are checked for validity, website building is tested, and format linting.
Cell Systems Article
To describe our effort, our training materials, and our philosophy, we wrote an article: Community-driven data analysis training for biology. We submitted the preprint on bioRxiv, got nice feedback, then submitted the full article on Cell Systems and got it accepted there with positive reviews!
So we are happy to say that the article has been published on the June issue!
An AWESOME Community!
This last year has been a success because of the community. Currently, >70 people have contributed at least once to the training material (>30 new contributors compared to last year). To highlight this awesome community, we have created a contributor Hall of Fame:
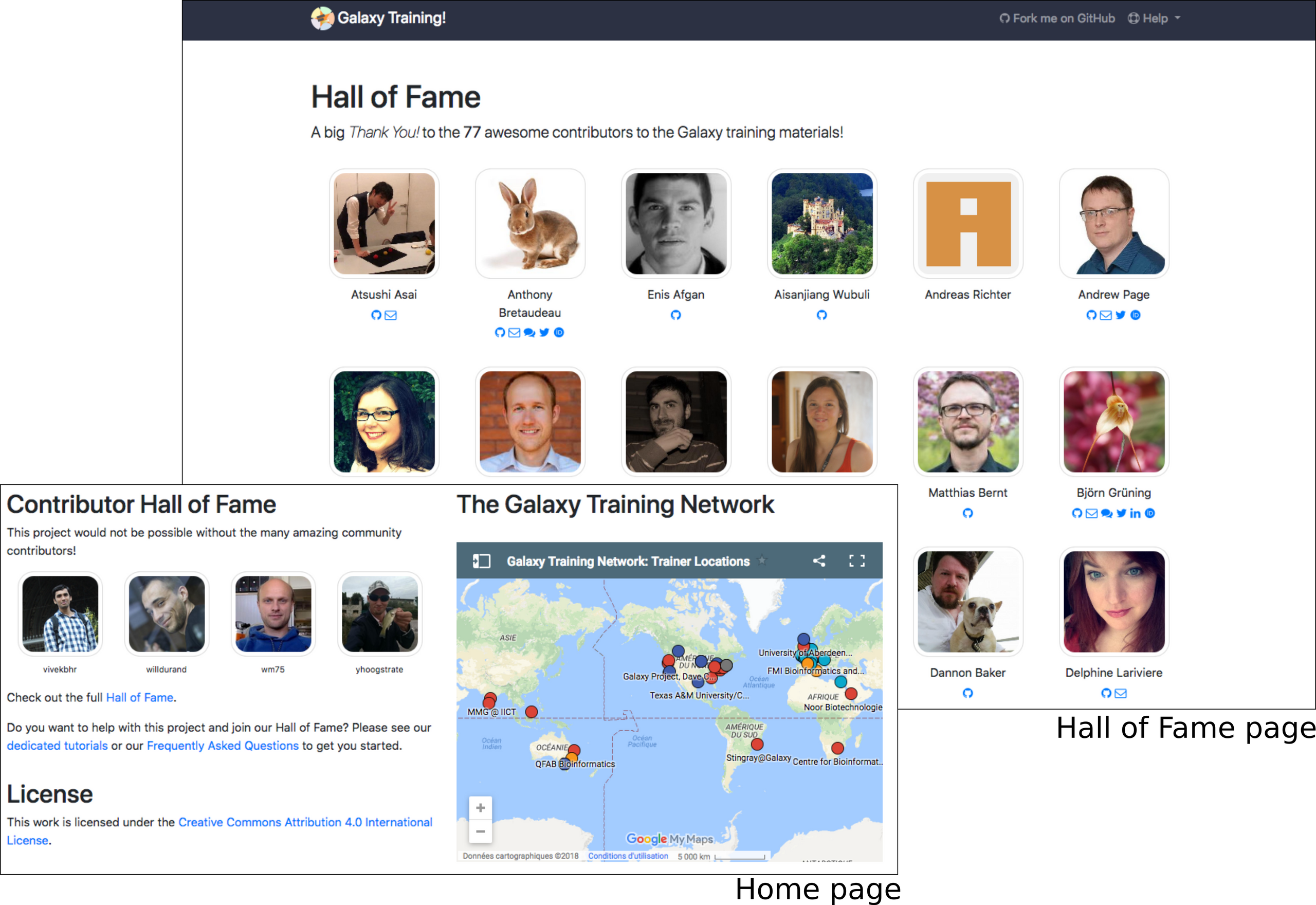
Each contributor is depicted by their GitHub profile image and can add information such as email, Gitter, Twitter, Linkedin, ORCID. Clicking on the images leads to the GitHub profile. To make this Hall of Fame visible on the homepage, we also added a little slide show on the homepage that cycles through all the contributors.
The new contributors are mostly joining during the community events, such as the Contribution Fests or Hackathons:

These events can be on-site, during conferencees or dedicated events; but we are also organizing online Contribution Fests (one or two days) to reach the whole community.
During these events, but also all over the year, our main communication channel continues to be our Gitter channel (read anonymously, but a GitHub or Twitter account is required to post) and our GitHub repository. We have labeled issues "newcomer-friendly" or "contribution fest") to help newcomers identify ways they can contribute.
As a community, we value diversity and inclusivity and we are eager to provide a welcoming and supportive environment for everyone, regardless of background or identity. We have added a Code of Conduct. We would like all interactions within our community to be underpinned by the Code of Conduct, whether it be contributions to GitHub or discussions on our Gitter channel. The Code is central to our efforts to build a welcoming, diverse, inclusive, and global community.
A Similarly Fruitful Future
This year has been quite full. Our infrastructure to deliver reproducible and robust training is now quite stable; we will be shifting our focus to the community.
For Users
We will continue to work on the content, expand it, and improve it. We plan to create small tutorials to explain some Galaxy tips, tutorials about visualization of data in Galaxy, etc. We would like to integrate more advanced tutorials on data analyses and create some topic-specific curriculum linking to the different tutorials. Some curriculum may include external references to R or Python tutorials, for example the R for genomics from Data Carpentry. This combination will be tested during workshops over the next year.
For Instructors
We have started to collect training best practices. We will continue this collection over the next year. We will also develop a mentorship program. Our idea would be to have a 1-hour online meeting once per month. During these meetings, we will discuss, exchange advice, receive feedback about the Galaxy training from the instructors. We would also like to develop some "Train the trainers" training, in collaboration with the ELIXIR Training Platform and The Carpentries.
For Contributors
We would like to improve the experience. Given all our effort, we know that contributing to the Galaxy Training material may be sometimes cumbersome and not always easy. We have started to implement some scripts to help create the skeleton of a tutorial, to automatically fill out the data-library.yaml and tools.yaml files, etc. We also developed a FAQ page and have added more tutorials to help contributing. We know that we can still do more; but first we need to identify the issues. During the next year, we will discuss together in person (during a BoF at GCC, during our different Contribution Fests, etc.) and collect feedback via a survey. Next year we hope to be able to show all of the results of these efforts.
For Everyone
We will organize more regular Contribution Fests. We plan to have an online Contribution Fest every 3 months on the 3rd Friday. The first one will be on the 17th of August where we will focus on metagenomics training; but anyone who would like to contribute on any other topics is very welcome to join. During these Contribution Fests, we will coordinate via Gitter and GitHub. We will also join bigger Contribution Fests, such as the GCC OBF Contribution Fest in Portland and the BioHackathon in Paris.
The next year will be hopefully as fruitful as this year! Thanks for the community for their investment in this nice project!
The statistics for the content and the contributors has been extracted using GitHub API. The scripts are available at: https://github.com/bebatut/galaxy-training-material-stats