February 2018 Galaxy News
On this page
Welcome to the February 2018 Galactic News, a summary of what is going on in the Galaxy community. If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events
There are a plenitude of Galaxy related events coming up in the next few months, including large regional meetings in Europe and Africa.
GCCBOSC 2018 Training Topic Voting Closes Jan 31
Which is tomorrow. Vote now
Your vote will determine the topics that are offered, which topics should be offered more than once, and which ones should not be scheduled at the same time. Your vote matters.
GCCBOSC2018 will be held 25-30 June in Portland, Oregon, United States. It will feature two days of training: the second of which is multi-track and will feature content for both the BOSC and Galaxy communities. Workshops will be hands-on and participants will be strongly encouraged to bring a laptop and follow along. If you work in data-intensive biomedical research, there is no better place than GCCBOSC 2018 to present your work and to learn from others.
Thanks to everyone who nominated topics and helped write topic descriptions!
GCCBOSC2018 registration will open in February. Watch this space.
GCCBOSC 2018 Sponsors
We are happy to have announced several GCCBOSC 2018 sponsors in January. This month we are highlighting PeerJ:
PeerJ
PeerJ is proud to sponsor GCCBOSC 2018 and support the unified event for bioinformatics developers and practitioners. PeerJ’s mission is to make Open Access affordable, fast, and easier for researchers and institutions, and central to this mission is our open source ethos. We publish two peer-reviewed journals, PeerJ and PeerJ Computer Science and we also host a free PeerJ Preprints server. PeerJ requires code to be open and accessible, hence readers can be assured that they can access the tools appropriately. We also provide open source tools helpful to both academics and general software development professionals.
GCCBOSC attendees may be interested to hear that, in addition to research articles, PeerJ welcomes articles describing bioinformatics software tools. These articles provide descriptions of new software tools (or significant new functionality in existing software) of interest to the bioinformatics and/or computational biology communities. You can browse the PeerJ Bioinformatics Software Tools Collection. Submit your bioinformatics manuscript here.
PeerJ is also excited to offer APC discounts to all GCCBOSC attendees - we’ll be in touch with more details soon!
[
 ](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_summary_20170906.pdf) [
](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_summary_20170906.pdf) [ ](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_prospectus_20170906.pdf)
](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_prospectus_20170906.pdf) Call for sponsors!
Sponsors are a key part of GCCBOSC 2018. Is your organization interested in playing a prominent role in the first joint gathering of the Galaxy and BOSC communities? Then become a GCCBOSC 2018 sponsor and raise your organization’s visibility in these active and engaged communities.
ELIXIR Galaxy Community Kickoff and Meeting, March, Freiburg
 ](https://www.elixir-europe.org/events/galaxy-community-kickoff-meeting-and-galaxy-user-conference)
](https://www.elixir-europe.org/events/galaxy-community-kickoff-meeting-and-galaxy-user-conference) [
 ](http://www.emblaustralia.org/)
](http://www.emblaustralia.org/)
The ELIXIR Galaxy community will have a kickoff meeting 14-16th March 2018, in Freiburg/Germany. We will combine this with a Galaxy User Conference and the official launch of the usegalaxy.eu server.
This event will also be coordinated with the EMBL Australia to also launch usegalaxy.org.au at the same time.
Wednesday (the 14th) will be dedicated to the ELIXIR Galaxy communtiy, to discuss and plan our roadmap in 2018 and 2019. Thursday and Friday (15th, 16th) will be a conference where Galaxy users talk about their research and use-cases. However, a few spots we will reserved for official talks from ELIXIR.
Register now as space is limited.
Galaxy Africa, April, Cape Town
Galaxy Africa will be held 3-5 April in Cape Town, South Africa at SANBI on the Universtiy of the Western Cape campus.
This is an opportunity to learn from leading bioinformaticists, systems administrators and engineers about Galaxy and accessible, reproducible analysis of biological data.
- Analyse: Learn about using Galaxy to analyse your genomic and RNA data.
- Manage: Meet fellow admins of research computing infrastructure and hear about local and cloud hosting of Galaxy servers.
- Extend: Hear how bioinformaticts are extending Galaxy through custom tools and workflows.
Our programme will start with keynote speakers introducing biology, data science and Galaxy. In the afternoon there will be tutorials on biological data analysis using Galaxy. The Wednesday morning session will include presentations on genomic and metagenomic data analysis workflows and uses of Galaxy. In the afternoon we will continue our tutorial series. The final day of the meeting will focus on the technology underlying Galaxy and biological data analysis with a focus on containerisation, software packaging for easier use and cloud deployment. Tutorials on this day will focus on similar topics.
Galaxy Administrators Course Report
 [
[ ](https://github.com/galaxyproject/dagobah-training)
](https://github.com/galaxyproject/dagobah-training)
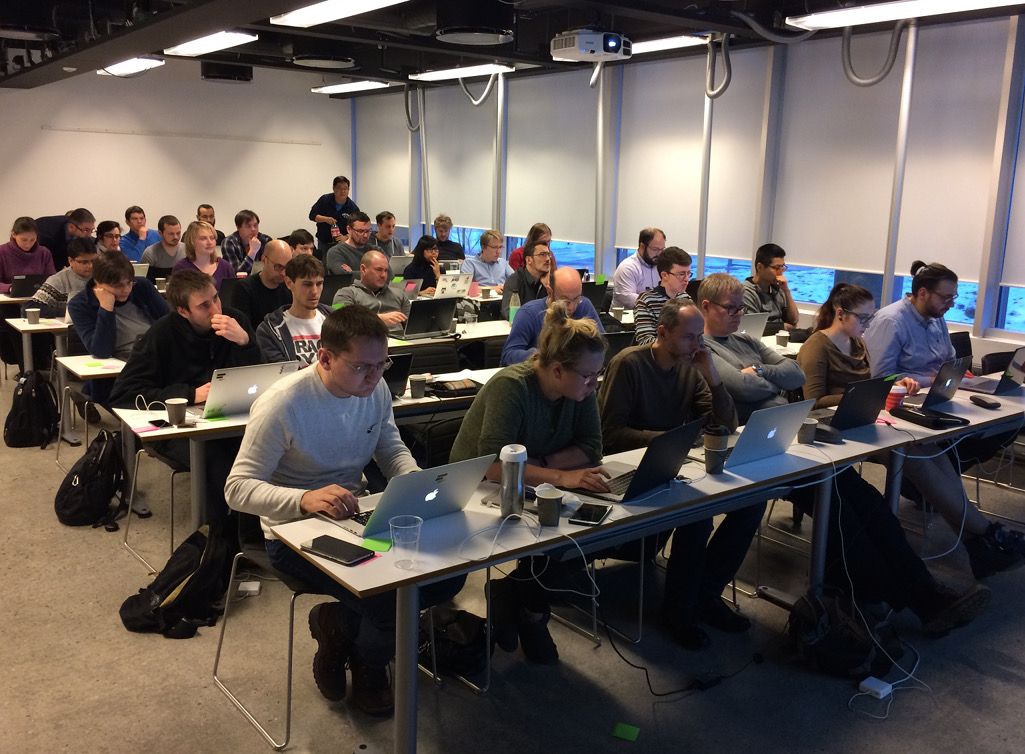
A very successful and inspiring Galaxy admin workshop was held in Oslo in January, Thanks to 33 highly motivated participants and Nicola Soranzo, Marius Van Den Beek, Abdulrahman Azab, Björn Grüning and Enis Afgan as instructors! Thanks also to ELIXIR, de.NBI, and ELIXIR Norway for sponsoring this event.
Plenty of materials for Galaxy administration are now up to date and available online. The workshop included a sneak preview of what’s coming in the 18.01 release from Nate Coraor.
Select Upcoming events
New Galactic Blog Entry: Galaxy R Markdown Tools
 ](/news/)
](/news/)
January saw one new Galactic Blog entry:
- Galaxy R Markdown Tools, by Ming Chen
Galaxy Cloud News
January was a big month for Galaxy in the cloud:
Publications
111 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in January.
Highlighted Publications
The Galactic and Stellar publications added in January were:
- META-pipe cloud setup and execution, Agafonov A, Mattila K, Tuan CD et al. F1000Research 2018, 6(ELIXIR):2060 (doi: 10.12688/f1000research.13204.2)
- Precise temporal regulation of alternative splicing during neural development, Sebastien M Weyn-Vanhentenryck, Huijuan Feng, Dmytro Ustianenko, Rachel Duffie, Qinghong Yan, Martin Jacko, Jose C Martinez, Marianne Goodwin, Xuegong Zhang, Ulrich Hengst, Stavros Lomvardas, Maurice S Swanson, Chaolin Zhang. bioRxiv 247601; doi: 10.1101/247601
- A-GAME: improving the assembly of pooled functional metagenomics sequence data, Matteo Chiara, Antonio Placido, Ernesto Picardi, Luigi Ruggiero Ceci, David Stephen Horner and Graziano Pesole, BMC Genomics 2018 19:44 doi: 10.1186/s12864-017-4369-z
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 67 | +Methods | 30 | +Workbench | 22 | +UsePublic | 10 | +UseMain | |||
| 8 | +UseLocal | 4 | +Reproducibility | 4 | +IsGalaxy | 3 | +RefPublic | |||
| 3 | +Tools | 3 | +Shared | 2 | +Other | 1 | +Visualization | |||
| 1 | +HowTo | 1 | +Unknown |
Who’s Hiring

The Galaxy is expanding! Please help it grow.
- The Freiburg Galaxy Team is looking for 2 Postdoctoral researchers
- The Blankenberg Lab in the Genomic Medicine Institute at the Cleveland Clinic Lerner Research Institute is hiring postdocs.
- Galaxy Project is hiring software engineers and postdocs at Johns Hopkins, Baltimore, Maryland, United States
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Public Galaxy Server News
There are over 90 publicly accessible Galaxy servers and six semi-public Galaxy services. Here’s what happened with them last month, including two new public servers:
A-Game
A-Game is A Galaxy suite for tArgeted MEtagenomics, a web service incorporating state of the art tools and workflows for the analysis of eDNA sequence data. A site manual and example workflows are available.
A-Game is supported by BEACON at the Dipartimento di Bioscienze, University of Milan
Splicescope
Splicescope is a tool-publishing server for predicting neuronal maturation using splicing profiles. Splicescope takes either an exon inclusion ratio matrix file or junction bed files from multiple samples as input and outputs a zip file including prediction results, PCA analysis results and a PCA plot based on reference samples and an html file to summarize the prediction.
Splicescope can be accessed anonymously, or with an account, and anyone can create an account. It is supported by the Zhang Laboratory, Department of Systems Biology, the Department of Biochemistry and Molecular Biophysics, and the Motor Neuron Center at Columbia University Medical Center.
New video from the Galaxy-P team
Public Servers in Publications
We tag papers that use, mention, implement or extend public Galaxy Servers. Here are the counts for January’s publications.
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 12 | >Huttenhower | 3 | >Workflow4Metabolomics | 1 | >GVL-Unspecified | 1 | >Galaxy-P | |||
| 1 | >Osiris | 1 | >PreSTIGE | 1 | >PIA | 1 | >Splicescope | |||
| 1 | >A-Game | 1 | >Cistrome | 1 | >Orione | 1 | >GVL-QLD | |||
| 1 | >RepeatExplorer |
Tools
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in January.
Releases
Earlier Releases
17.09 Galaxy Release
The Galaxy Committers published the 17.09 release of Galaxy at the end of October.
Highlights include
- Singularity Tool execution using the HPC-friendly container technology Singularity is now supported.
- Download entire collection Downloading whole collections is now possible from the history interface. (Thanks to @mvdbeek.)
- Switch tool versions in workflows You can now select exactly what version of tool you want to use when building workflows. (Thanks to @mvdbeek.)
- Security patches
- Several features were deprecated
See the 17.09 release announcement for details.
Galaxy Docker Image 17.09
The Galaxy Docker project has seen a matching release, for Galaxy 17.05. Major features include
- much improved documentation about using Galaxy Docker and an external cluster (@rhpvorderman)
- CVMFS support - mounting in 4TB of pre-build reference data (@chambm)
- Singularity support and tests (compose only)
- more work on K8s support and testing (@jmchilton)
- using .env files to configure the compose setup for SLURM, Condor, K8s, SLURM-Singularity, Condor-Docker
And
- The Galaxy Docker Project has reached more than 31k downloads on Dockerhub - not counting quay.io and all flavors
ephemeris 0.8.0
Ephemeris is a small Python library and set of scripts for managing the bootstrapping of Galaxy plugins - tools, index data, and workflows.
- Free software: Academic Free License version 3.0
- Documentation: https://ephemeris.readthedocs.org.
- Code: https://github.com/galaxyproject/ephemeris
blend4php 0.1 beta
The beta version of the blend4php package, a PHP wrapper for the Galaxy API, was released in December. It provides a PHP package for interacting with Galaxy and CloudMan. blend4php currently offers a partial implementation of the Galaxy API and includes support for datasets, data types, folder contents, folders, genomes, group roles, groups, group users, histories, history contents, jobs, libraries, library contents, requests, roles, search, tools, toolshed repositories, users, visualizations and workflows.
The motivation for development of this library is for integration with Tripal, an open-source toolkit for creation of online genomic, genetic and biological databases.
Please see the API documentation page for full information.
galaxy-lib 17.9.10
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
This revision:
- Added docs for using mulled-build with your own quay.io account (thanks to @jerowe).
- Catch errors in Conda search if nothing is found (preventing planemo-monitor from functioning properly) (thanks to @bgruening).
- Make multi-requirement container building via mulled more stable (thanks to @bgruening).
Planemo 0.47.0
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. These releases included numerous fixes and enhancements.
See GitHub for details.
Other packages that have been released in the prior 4 months.
sequence_utils 1.1.2
Galaxy’s sequence utilities are a set of Python modules for reading, analyzing, and converting sequence formats. See the release notes for what’s new this month.
StarForge 0.3.1-5
StarForge help build Galaxy things in virtualization:
- Build Galaxy Tool Shed dependencies
- Build Python Wheels (e.g. for the Galaxy Wheels Server)
- Rebuild Debian or Ubuntu source packages (for modifications)
These things will be built in Docker. Additionally, wheels can be built in QEMU/KVM virtualized systems. StarForge has had several updates this fall. Fixes and new features include:
- Support xz/lzma tarballs for wheel builds Pull Request 166
- Native support for auditwheel and delocate. (#160)
- Do not build sdists with the wheel subcommand by default. (#155)
- Fix a bug where the wrong working directory was set when building wheels with multiple sources. (#154)
- Fix a bug with sudo and brew install on macOS. (#151).
- Short circuit platform caching on OS X (#150).
And the rest …
Other Galaxy packages that haven’t had a release in the past four months can be found on GitHub.
Other News
- The semi-annual update of the project statistics page is done. See what’s new.
- From John Chilton:
- Galaxy can now by-pass re-running jobs with duplicate parameters/inputs thanks to the brilliant Marius Van Den Beek