February 2016 Galaxy News
On this page
Welcome to the February 2016 Galactic News, a summary of what is going on in the Galaxy community.
If you have anything to include in the next News, please send it to [Galaxy Outreach](mailto:outreach AT galaxyproject DOT org).
GCC2016
GCC2016 will be held June 25-29 at Indiana University in Bloomington, Indiana, United States. This will be the 7th annual gathering of the Galaxy community, and we are expecting over 200 participants again this year. The 2016 Galaxy Community Conference includes 2 days of hackathons, 2 days of training, and a two day meeting featuring accepted presentations, keynotes, poster sessions, the new Visualization Showcase and Software Demo sessions, lightning talks, birds-of-a-feather meetups, and plenty of networking.
GCC2016 Early Registration is Open
We are pleased to announce that early registration for GCC2016 is now open. Registration costs depend on which events you register for, your career stage & affiliation, and when you register. Early bird registration ends May 20 and is up to 40% less than regular registration rates. Early bird registration starts at less than $45 / day for students and postdocs, and at $65 / day for other attendees from non-profits.
You can also sign up for conference housing during registration.
You are strongly encouraged to review the training and housing options before beginning the registration process.
GCC2016 Abstract Submission is Open
Abstract submission for GCC2016 is now open. We are accepting oral and poster presentation abstracts, as well as proposals for the brand new Visualization Showcase and Software Demo track. You may submit similar content for oral presentations, posters, and demos. Topics should be of interest to those working in high-throughput data analysis and research. Presentations that are Galaxy-centric are encouraged, but not required. If you are submitting a demo you are encouraged to consider submitting a corresponding poster.
The deadline for oral presentation abstracts is March 25, 2016. Abstracts will be reviewed and submitters will be notified of acceptance status no later than April 11, 2016.
Poster and Demo abstract submission closes May 20, 2016 (or earlier depending on poster space availability). Poster and demo submissions are reviewed on a rolling basis, and submitters will be notified of acceptance status no later than two weeks after the abstract is submitted. Please see the abstracts page for full details.
This is an ideal time to share your work with the community.
Sponsors
And we are pleased to welcome Lenovo, as a first time Gold Sponsor of GCC. Lenovo offers a comprehensive portfolio of high-performance computing infrastructure designed to accelerate research, reduce IT complexity and maximize asset utilization in the life sciences. Lenovo and its network of partners offer a broad portfolio of open, industry-standard infrastructure solutions that can help accelerate product innovation without becoming locked into expensive, inflexible proprietary environments. Lenovo Solution for Life Sciences is comprised of pre-integrated, high-performance servers, storage systems and networking equipment, advanced file systems and integrated workflow management software. Designed to accommodate large data volumes and compute-intensive applications, this solution can help organizations accelerate time to results at a lower total cost of ownership while making it easier to scale your HPC cluster as business grows.
Also, please welcome Globus Genomics as a GCC sponsor for the 4th year in a row. Globus Genomics is an integrated solution for Next Gen Sequencing analysis utilizing technologies in big data management and big data analysis. Globus Genomics combines big data management capabilities of Globus Online with flexible, intuitive workflow/pipeline creation capabilities of the Galaxy framework and high throughput computing capabilities on Amazon Web Services. Using Globus Genomics, researchers can easily transfer large amounts of sequence data from various sequencing centers and analyze the data either interactively or using one of the pre-defined best practice analytical pipelines with the familiar Galaxy interface.
We continue to seek other sponsors as well and [offer a wide range of sponsorship plans](https://web.archive.org/web/http://gcc2016.iu.edu/sponsors/index.php). If your organization is interested in having a presence at GCC2016, please contact the GCC2016 Exec for more information.
New Papers
156 new papers referencing, using, extending, and implementing Galaxy were added to the Galaxy CiteULike Group in January. (Note: that is a record, but it reflects back-curation efforts for papers from 2014 & 2015 that we couldn’t curate the first time.) Highlights include:
-
Varying levels of complexity in transcription factor binding motifs Jens Keilwagen, Jan Grau, Nucleic Acids Research, Vol. 43, No. 18. (26 June 2015), pp. gkv577-e119, doi:10.1093/nar/gkv577
-
Scaling Up Bioinformatics Workflows with Dynamic Job Expansion: A Case Study Using Galaxy and Makeflow Nicholas Hazekamp, Joseph Sarro, Olivia Choudhury, Sandra Gesing, Scott Emrich, Douglas Thain, In e-Science (e-Science), 2015 IEEE 11th International Conference on (August 2015), pp. 332-341, doi:10.1109/escience.2015.39
-
RiboTools: a Galaxy toolbox for qualitative ribosome profiling analysis Rachel Legendre, Agnès Baudin-Baillieu, Isabelle Hatin, Olivier Namy, Bioinformatics, Vol. 31, No. 15. (01 August 2015), pp. 2586-2588, doi:10.1093/bioinformatics/btv174
-
Integrating UIMA with Alveo, a human communication science virtual laboratory Dominique Estival, Steve Cassidy, Karin Verspoor, Andrew MacKinlay, Denis Burnham, In Proceedings of the Workshop on Open Infrastructures and Analysis Frameworks for HLT (August 2014), pp. 12-22, doi:10.3115/v1/W14-5202
-
Alveo, a Human Communication Science Virtual Laboratory Dominique Estival, Steve Cassidy, In Proceedings of the Australasian Language Technology Association Workshop 2014 (November 2014), pp. 104-107
-
Integrating Data-Intensive Computing Systems with Biological Data Analysis Frameworks Edvard Pedersen, Inge A. Raknes, Martin Ernstsen, Lars Ailo Bongo, In Parallel, Distributed and Network-Based Processing (PDP), 2015 23rd Euromicro International Conference on (March 2015), pp. 733-740, doi:10.1109/pdp.2015.106
-
Deciphering regulation in eukaryotic cell: from sequence to function Valentina Boeva, (23 Sep 2014)
-
Galaxy Portal: Interacting with the Galaxy platform through mobile devices Claus Børnich, Ivar Grytten, Eivind Hovig, Jonas Paulsen, Martin Čech, and Geir Kjetil Sandve, Bioinformatics first published online January 27, 2016 doi:10.1093/bioinformatics/btw042
The new papers were tagged with:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 10 | Cloud | 13 | Other | - | Shared | 17 | UseMain | |||
| 1 | HowTo | - | Project | 8 | Tools | 12 | UsePublic | |||
| 4 | IsGalaxy | 9 | RefPublic | - | UseCloud | 3 | Visualization | |||
| 56 | Methods | 2 | Reproducibility | 8 | UseLocal | 57 | Workbench |
Events
Upcoming Events
There are many upcoming events in the next few months. See the Galaxy Events Google Calendar for details on other events of interest to the community.
| |
Designates a training event offered by GTN member(s) |
Who’s Hiring
The Galaxy is expanding! Please help it grow.
- Postdoc position on methods for transcriptome analysis of disease-specific cells, University of Oslo
- Computational Biology post-doc, IMI-eTRIKS consortium, CNRS-EISBM, Lyon France. Contribute “to the development of eTRIKS Galaxy tools and workflows for disease stratification and biomarker discovery from single and multiple omics datasets.”
- Post-doc in Functional Genomics Of Obesity-Related Diseases, Inserm, Lille, France
- Software Engineer, Oregon Health Sciences University, Portland, Oregon, United States
- The Galaxy Project is hiring software engineers and post-docs
Got a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
New Tutorials … and Tours!
A few new tutorials are now available on GitHub.
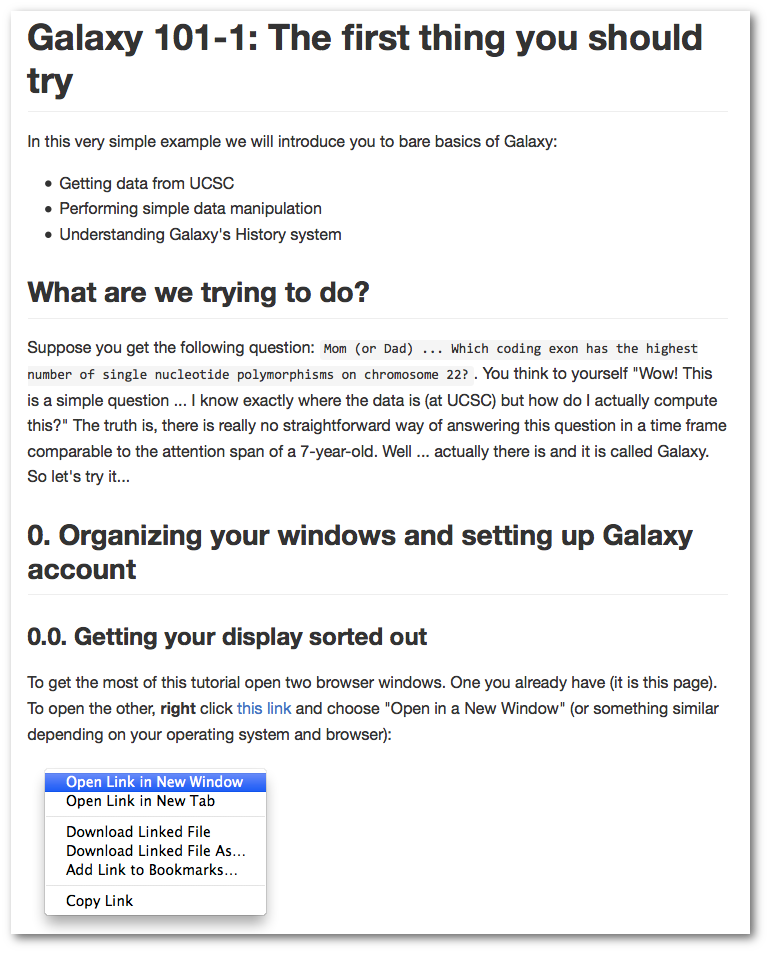
An updated and expanded Galaxy 101
First there is an updated and expanded version of the Galaxy 101 Tutorial. Galaxy 101 introduces the Galaxy platform, how to get data from UCSC, and performing data analysis using genomic coordinates in multiple datasets. The tutorial also covers the basics of Galaxy Histories, creating and using re-runnable workflows.
Processing many samples at once
There is a new tutorial on using dataset collections titled Processing many samples at once. From the tutorial
Here we will show Galaxy features designed to help with the analysis of large numbers of samples. When you have just a few samples - clicking through them is easy. But once you’ve got hundreds - it becomes very annoying. In Galaxy we have introduced Dataset collections that allow you to combine numerous datasets in a single entity that can be easily manipulated.
Running Du Novo interactively from Galaxy: An ABL1 example
And there is also a new tutorial, Running Du Novo interactively from Galaxy: An ABL1 example, that “explains how to perform discovery of low frequency variants from duplex sequencing data” using an example ABL1 dataset from Schmitt and colleagues (SRA accession SRR1799908).
Galaxy Interactive Tours
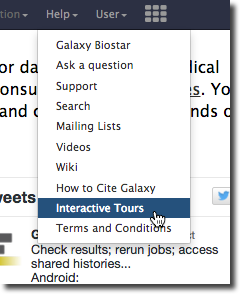
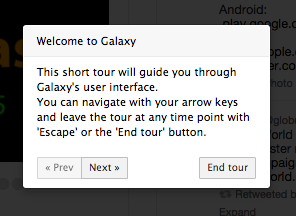
UseGalaxy.org now supports interactive guided tours and two introductory tours are available:
Interactive tours are a way to walk through Galaxy, following a set of steps to accomplish a task or learn a feature. The list of supported interactive tours on any server can be found in the Help menu. Support for interactive tours will be included in the 16.01 Galaxy release. Look for more tours in future releases.
----
Galaxy on Windows and OS X using Kitematic
Galaxy has been available in Docker containers for a while now. Docker containers are an easy way to package software for installation on other systems. The Docker Toolbox now includes Kitematic, a GUI for running Docker containers on Windows and Mac OS X systems. Kitematic makes it easy to run any published Docker container on these systems.
Want to try a pre-configured Galaxy instance on your Mac OS X or Windows machine? Try these steps:
- Install the Docker Toolbox on your computer.
- Launch Kitematic.
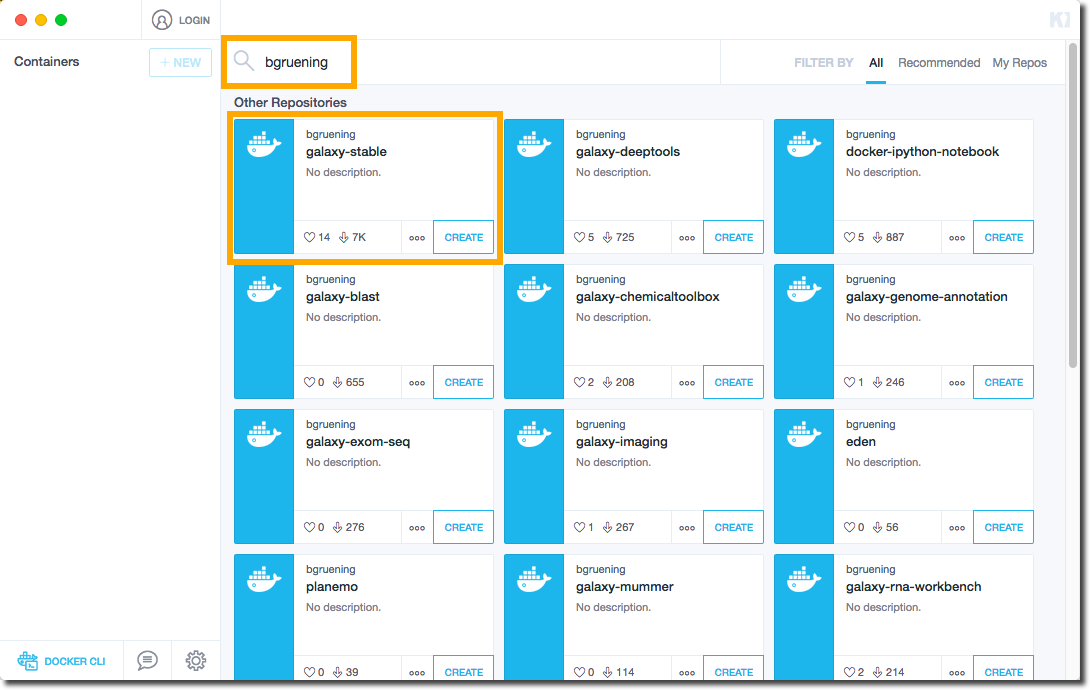
- Search for “galaxy” and select one of the relevant Galaxy options. This searches Docker Hub, a repository for Docker containers. “bgruening/galaxy-stable” will launch the latest generic Galaxy instance.
- Once the instance has started, click on the web preview pane, and voilà, you have a running Galaxy instance.
New Public Galaxy Servers
One new publicly accessible Galaxy servers was added in January: LAPPS Grid, a Galaxy server for natural language processing.
LAPPS Grid
* *Links:* * [LAPPS Grid Galaxy Workflow Engine](http://galaxy.lappsgrid.org/), part of the [Languarge Application Grid](http://www.lappsgrid.org/)- Domain/Purpose:
- “An open, interoperable web service platform for natural language processing (NLP) research and development. The LAPPS Grid provides facilities to select from hundreds of NLP tools to create workflows, composite services, and applications, and to evaluate, reproduce, and share them with others.”
- Comments:
- “This is a Work In Progress. Many of the services here are undergoing active development and their behaviour is likely to change without notice.”
- User Support:
- Landing page includes a simple tutorial.
- Quotas:
- Anyone can create a login, or it can be used anonymously
- Sponsor(s):
- See the LAPPS Grid Participants page. Includes
Releases
galaxy-lib 16.1.8 - 16.1.9
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
Starforge 0.1
Starforge is a collection of scripts that supports the building of components for Galaxy. Specifically, with Starforge you can:
- Build Galaxy Tool Shed dependencies
- Build Python Wheels (e.g. for the Galaxy Wheels Server)
- Rebuild Debian or Ubuntu source packages (for modifications)
These things will be built in Docker. Additionally, wheels can be built in QEMU/KVM virtualized systems.
Documentation can be found at starforge.readthedocs.org.
Other Releases
Galaxy 15.10
The October 2015 Galaxy Release (v 15.10) was released on November 30. The Reports Application, data upload widget, and data libraries all saw significant work.
See the full release notes for details.
Galaxy Docker Image 15.10
And, thanks to Björn Grüning, there is also now a Docker image for Galaxy 15.10 as well.
CloudMan 15.12
CloudMan is a cloud manager that orchestrates all of the steps required to provision a complete compute cluster environment on a cloud infrastructure; subsequently, it allows one to manage the cluster, all through a web browser.
This minor CloudMan 15.12 release includes:
- Update Galaxy to the Galaxy 15.10 release (see above for all the juicy details with the improvements that brings along)
- Preload the GIE IPython Docker container onto the image for faster startup: less than 30 seconds to a fully functional IPython notebook vs. several minutes before!
- For those of you that ran into the problem of the size of your user data overfilling after having added and removed a bunch of worker instances - no worries, that has been fixed now and resource tags are no longer stored in the cluster config to prevent persistent_data.yaml from becoming too big.
- The file system archive extraction has been made more resilient and synchronous.
- For those that are building custom images, a more direct method for locating the Nginx configuration directory has been added to help with different versions of the operating system (thanks to Matthew Ralston)
- A number of dependent library versions have been updated to their latest versions.
Pulsar 0.6.1
Pulsar 0.6 was released in December. Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
The 0.6.x release includes these changes:
- Pulsar now depends on the new galaxy-lib Python package instead of manually synchronizing Python files across Pulsar and Galaxy.
- Numerous build and testing improvements.
- Fixed a documentation bug in the code (thanks to @erasche). e8814ae
- Remove galaxy.eggs stuff from Pulsar client (thanks to @natefoo). 00197f2
- Add new logo to README (thanks to @martenson). abbba40
- Implement an optional awknowledgement system on top of the message queue system (thanks to @natefoo). Pull Request 82 431088c
- Documentation fixes thanks to @remimarenco. Pull Request 78, Pull Request 80
- Fix project script bug introduced this cycle (thanks to @nsoranzo). 140a069
- Fix config.py on Windows (thanks to @ssorgatem). Pull Request 84
- Add a job manager for XSEDE jobs (thanks to @natefoo). 1017bc5
- Fix pip dependency installation (thanks to @afgane) Pull Request 73
Planemo 0.21.1
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. Planemo 0.21.1 is the most recent release. See the release history.
BioBlend 0.7.0
BioBlend version 0.7.0 was released at the beginning of November. BioBlend is a python library for interacting with CloudMan and the Galaxy API. CloudMan offers an easy way to get a personal and completely functional instance of Galaxy in the cloud in just a few minutes, without any manual configuration.) From the release CHANGELOG.
blend4j v0.1.2 blend4j v0.1.2 was released in December 2014. blend4j is a JVM partial reimplemenation of the Python library bioblend for interacting with Galaxy, CloudMan, and BioCloudCentral.
Galaxy Community Hubs
 |
 |
 |
| Share your training resources and experience now | Share your experience now |
ToolShed Contributions
See list of tools contributed in January.
Other News
- The Project Statistics page has had its semi-annual update.
Galaxy Goes Mobile!
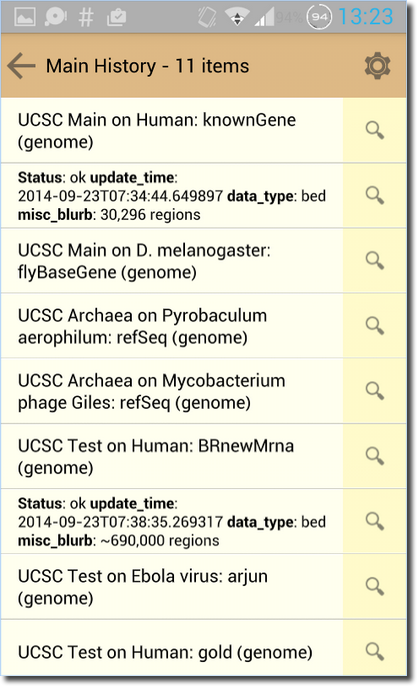
Galaxy is now accessible from your IOS and Android devices. The Galaxy Portal app is a quick and easy way to monitor the status of biomedical research on any Galaxy server. With this app you can set up a list of Galaxy connections and browse your analyses histories in a user-friendly format and take a peek at your data on the go.
The app has been in development for over a year at the University of Oslo as part of Claus Børnich’s masters thesis and is described in
Claus Børnich, Ivar Grytten, Eivind Hovig, Jonas Paulsen, Martin Čech, and Geir Kjetil Sandve. Galaxy Portal: Interacting with the Galaxy platform through mobile devices. Bioinformatics first published online January 27, 2016 doi:10.1093/bioinformatics/btw042
A user manual, additional information, and the full source are available on GitHub.
Galaxy Portal is available free of charge.