April 2018 Galaxy News
On this page
Welcome to the April 2018 Galactic News, a summary of what is going on in the Galaxy community. If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
UseGalaxy.eu and UseGalaxy.org.au Launched
UseGalaxy.org is absolutely delighted to announce the birth of two sibling servers - UseGalaxy.eu and UseGalaxy.org.au. The three UseGalaxy servers will mirror a set of core tools and reference genomes and are aimed at better distributing access to Galaxy around the world.
UseGalaxy.eu is a publicly accessible server that aims to support the analysis and sharing needs of the European research community. It was launched in March 2018 at European Galaxy User Meeting in Freiburg. It is hosted at the University of Freiburg. See:
- Green Light for Galaxy Europe
- Flyer with more information
- UseGalaxy.eu public server directory entry
UseGalaxy.org.au aims to support the Australian research community. It is built on the Genomics Virtual Laboratory infrastructure, and was also launched in March 2018. This server was formerly the University of Queensland Galaxy Server. See:
- Synergising Australian bioinformatics resources through the launch of Galaxy Australia
- Announcing an Australia-wide Galaxy service
- UseGalaxy.org.au public server directory entry
And see the new UseGalaxy page for future plans.
GCCBOSC 2018
GCCBOSC 2018 will be held 25-30 June in Portland, Oregon, United States. This brings the 2018 Galaxy Community Conference and the Bioinformatics Open Source Conference together into a unified week-long event. If you work in open source life science or data-intensive biomedical research, then there is no better place than GCCBOSC 2018 to present your work and to learn from others.
GCCBOSC starts with two days of training with a wide range of topics nominated and selected by our communities. Training is followed by a two day meeting, with joint and parallel tracks, featuring oral presentations, posters, demos, lightning talks, birds-of-a-feather and invited keynotes. The week finishes with CollaborationFest Core and Encore, two or four days of collaborative work on code, documentation, training and challenging data analysis problems.
Keynote speakers
We are pleased to announce our third keynote speaker:
GCC2018 Keynote: Lucia Peixoto
- Elson S. Floyd College of Medicine, Washington State University
- Keynote title: Confound it! Reproducible biology from “omics” data analysis
Dr. Peixoto is an Assistant Professor at the Elson S. Floyd College of Medicine at Washington State University. Her research focuses on using genomic and computational biology approaches to study brain function. She also advises the WSU Spokane Genomics Core, and has published work on how analysis of complex datasets affects power and reproducibly in RNA-seq and Epigenomic-seq.
Submit GCC lightning talks, posters and demos by April 16
The deadline for submitting lightning talks for GCC2018 is April 16, less than a week away. You are strongly encouraged to submit a talk (or two) by the deadline. Depending on response we may also open up for late-breaking lightning talks in May.
On-time poster and demo submissions are also closing April 16. To assure a place for your poster and/or demo, please submit these by that deadline as well. Depending on space, we will (likely) also open a call for late-breaking posters and demos in May.
Early registration ends May 11
So c’mon, register already:
Childcare at GCCBOSC
Subsidized childcare for children 6 weeks to 12 years old is available at Portland State University for $7-10 / hour. If you want, you can also bring your infant with you to the conference. A nursing room will be available on any day it is requested.
Interested? Please register your interest in a nursing room, and/or sign up for childcare now. Space in childcare is very limited.
GCCBOSC 2018 Sponsors
We are happy to have several confirmed GCCBOSC 2018 sponsors. This month we are highlighting Google Cloud, a GCCBOSC 2018 Platinum Sponsor:
Google Cloud
Google Cloud is working closely with the healthcare and life science industry to provide the technology and tools that power the innovations and discoveries that improve our healthcare system. By providing advanced, healthcare-specific technology solutions, we hope to accelerate our customers ability to change the way healthcare discoveries are made. Through data management at scale, advancements in machine learning, APIs, and on-demand infrastructure, we offer solutions that enable provider, research, and life science customers to tackle the rapidly evolving and long standing challenges facing the healthcare industry.
[
 ](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_summary_20170906.pdf) [
](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_summary_20170906.pdf) [ ](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_prospectus_20170906.pdf)
](https://depot.galaxyproject.org/hub/attachments/documents/gccbosc2018_sponsorship_prospectus_20170906.pdf) Call for sponsors!
Sponsors are a key part of GCCBOSC 2018. Is your organization interested in playing a prominent role in the first joint gathering of the Galaxy and BOSC communities? Then become a GCCBOSC 2018 sponsor and raise your organization’s visibility in these active and engaged communities.
2018 Big Genomics Data Skills Training course
There are still a few spaces left in Jackson Lab (JAX)‘s 2018 Big Genomics Data Skills Training course, funded by the NIH BD2K initiative: Big Genomic Skills Training info and online application. The workshop will enable participants to integrate genomic data analysis into their courses or launch new courses. Participants will gain experience using Galaxy, R and Python.
Housing is included; however you will need to cover your own travel. If you teach at an HBCU or Minority-serving institution (by Dept. of Education criteria) JAX may be able to provide a modest travel award.
The course is for faculty who primarily teach undergraduate students and will be held at JAX Genomic Medicine in Farmington CT, May 21-25.
Gateways 2018 Call for Participation (1st deadline: May 7)
Gateways 2018 (September 25–27, at the University of Texas at Austin) is now accepting submissions of papers, demos, tutorials, and panels on the topic of science or engineering gateways. Gateways are user-friendly interfaces (including Galaxy) to scientific computing, data, and other domain-specific resources to support research and education.
Topics may include their design, use, impact, development processes, sustainability, best practices, or any other aspect that you think fellow gateway creators or users will find interesting to learn. We also welcome educational topics directed toward the next generation of gateway creators.
The primary submission deadline is May 7, 2018, and a poster session deadline (open to all) will be August 1. Read more details in the Call for Participation
Upcoming events
These and other Galaxy related events coming up in the next few months:
Publications
119 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in March.
Highlighted Publications
March was a banner month for Galactic and Stellar publications:
- WAVES: a Web Application for Versatile Enhanced bioinformatic Services, Marc Chakiachvili, Anne-Muriel Arigon Chifolleau, Vincent Lefort. bioRxiv, doi: 10.1101/276485
- Using the FACE-IT portal and workflow engine for operational food quality prediction and assessment: An application to mussel farms monitoring in the Bay of Napoli, Italy, Raffaele Montella, AlisonBrizius, Diana Di Luccio, Cheryl Porter, Joshua Elliot, Ravi Madduri, David Kelly, Angelo Riccio, Ian Foster. Future Generation Computer Systems doi: 10.1016/j.future.2018.03.002
- ImmPort, toward repurposing of open access immunological assay data for translational and clinical research, Sanchita Bhattacharya, Patrick Dunn, Cristel G. Thomas, Barry Smith, Henry Schaefer, Jieming Chen, Zicheng Hu, Kelly A. Zalocusky, Ravi D. Shankar, Shai S. Shen-Orr, Elizabeth Thomson, Jeffrey Wiser & Atul J. Butte. Scientific Data volume 5, Article number: 180015 (2018), doi:10.1038/sdata.2018.15
- SMAGEXP: a galaxy tool suite for transcriptomics data meta-analysis, Samuel Blanck, Guillemette Marot. arXiv:1802.08251 [q-bio.GN]
- META-pipe – Distributed Pipeline Analysis of Marine Metagenomic Sequence Data, PhD. Thesis, Espen Mikal Robertsen
- Discovery of coding regions in the human genome by integrated proteogenomics analysis workflow, Yafeng Zhu, Lukas M. Orre, Henrik J. Johansson, Mikael Huss, Jorrit Boekel, Mattias Vesterlund, Alejandro Fernandez-Woodbridge, Rui M. M. Branca & Janne Lehtiö. Nature Communications volume 9, Article number: 903 (2018) doi:10.1038/s41467-018-03311-y
- Development and evaluation of a culture-free microbiota profiling platform (MYcrobiota) for clinical diagnostics, Stefan A. Boers, Saskia D. Hiltemann, Andrew P. Stubbs, Ruud Jansen, John P. Hays. European Journal of Clinical Microbiology & Infectious Diseases (2018), doi: 10.1007/s10096-018-3220-z
- A Distinctive Urinary Metabolomic Fingerprint Is Linked With Endoscopic Postoperative Disease Recurrence in Crohn’s Disease Patients, Ammar Hassanzadeh Keshteli, Robert Tso, Levinus A Dieleman, Heekuk Park, Karen I Kroeker, Juan Jovel, Patrick M Gillevet, Masoumeh Sikaroodi, Rupasri Mandal, Richard N Fedorak, Karen L Madsen. Inflammatory Bowel Diseases, Volume 24, Issue 4, 19 March 2018, Pages 861–870, doi: 10.1093/ibd/izx070
- Adapting the Smart-seq2 Protocol for Robust Single Worm RNA-seq, Lorrayne Serra, Dennis Chang, Marissa Macchietto, Katherine Williams, Rabi Murad, Dihong Lu, Adler Dillman, Ali Mortazavi. Bio-Protocol doi: 10.21769/BioProtoc.2729
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 66 | +Methods | 27 | +Workbench | 22 | +UsePublic | 13 | +UseMain | |||
| 8 | +RefPublic | 8 | +UseLocal | 7 | +Tools | 7 | +Reproducibility | |||
| 5 | +IsGalaxy | 3 | +HowTo | 3 | +Shared | 3 | +Cloud | |||
| 2 | +Unknown | 1 | +Other |
Who’s Hiring

The Galaxy is expanding! Please help it grow.
- System developers / bioinformaticians, two positions, ELIXIR Norway, University of Oslo, Norway (closes 17 April)
- Scientist (f/m), NGS bioinformatics core facility,Helmholtz Zentrum München, Germany.
- PhD position: Proteome Biology of Solid Tumors: Targeted, Spatially Resolved & Explorative Approaches, University of Freiburg, Germany
- Ingenieur Bioinformatique Evolutive LIRMM, Montpellier, France
- Post-Doc / IR analyse de données miRNA, Unité de Nutrition Humaine, Clermont-Fd, France
- Freiburg Galaxy Team has open positions, Freiburg, Germany
- Development of innovative algorithms to process real-time mass spectrometry data for the clinical analysis of exhaled breath, CEA, Saclay, France
- The Blankenberg Lab in the Genomic Medicine Institute at the Cleveland Clinic Lerner Research Institute is hiring postdocs.
- Galaxy Project is hiring software engineers and postdocs at Johns Hopkins, Baltimore, Maryland, United States
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Public Galaxy Server News
There are over 100 publicly accessible Galaxy servers and six semi-public Galaxy services. In addition to the UseGalaxy launches, another public server was added to the directory, and one was updated:
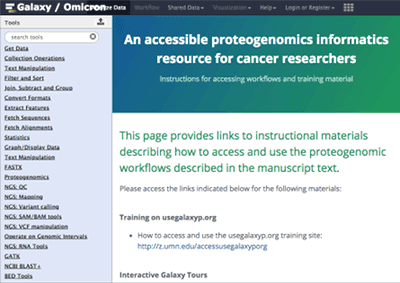
Proteogenomics Gateway
The Galaxy-P Proteogenomics Gateway provides access to documentation and other instructional materials, and an opportunity for hands-on training using example datasets and optimized proteogenomics workflows. The main goal of this server is to provide documentation to facilitate training and mastery of these software and workflows.
Step-by-step training instructions and a wealth of other material are available. The server can be accessed anonymously or you can create an account. This Galaxy is also available on Jetstream. It is maintained by @galaxyproteomics.
See An Accessible Proteogenomics Informatics Resource for Cancer Researchers by Matt Chambers et al. for more.
PhenoMeNal release Cerebellin now live
We are pleased to announce the 2018.02 release “Cerebellin” of the PhenoMeNal infrastructure. The new release is the result of half a year of intense developments and represents a major upgrade to the 2017-08 production release Bucetin. The main features of this release are:
- Improved usability following extensive User experience tests
- The Infrastructure has improved stability and resilience, through optimised configuration and resource allocation
- Microsoft Azure as new cloud target
- New ISA-Tab datatype in Galaxy
- The first interactive Galaxy tours have been added that help follow the steps in a workflow
For more information about the release, see Release notes 2018 02.
Public Servers in Publications
We tag papers that use, mention, implement or extend public Galaxy Servers. Here are the counts for March’s publications.
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 9 | >Huttenhower | 2 | >DeepTools | 2 | >Workflow4Metabolomics | 2 | >ErasmusMC | |||
| 2 | >GVL-Cloud | 2 | >Mississippi | 1 | >Cistrome | 1 | >EuPathDB | |||
| 1 | >ARGs-OAP | 1 | >ImmPort | 1 | >MPI-HLR | 1 | [>Genomic Hyperbrowser](https://www.zotero.org/groups/1732893/galaxy/tags/>Genomic Hyperbrowser) | |||
| 1 | >SCDE | 1 | >RepeatExplorer | 1 | >ARGalaxy | 1 | [>In Silico](https://www.zotero.org/groups/1732893/galaxy/tags/>In Silico) | |||
| 1 | >SouthGreen | 1 | [>LAPPS Grid](https://www.zotero.org/groups/1732893/galaxy/tags/>LAPPS Grid) | 1 | >VarCap | 1 | [>RNA Workbench](https://www.zotero.org/groups/1732893/galaxy/tags/>RNA Workbench) | |||
| 1 | >PhenoMeNal | 1 | >SSC | 1 | >SMAGEXP | 1 | >PreSTIGE | |||
| 1 | >Galaxy-P |
Tools
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in March.
Releases
18.01 Galaxy Release
The 18.01 release of Galaxy is out. Thanks to the Galaxy Committers and you, our community!
Highlights:
Performance and User Experience Improvements We made Galaxy more lively and responsive. Homepage, published workflows, published/saved histories, and data libraries should all load much faster now. Importing data from FTP will also take less of your time.
Web Server and Configuration The default web server used by Galaxy has changed from Paste to uWSGI and the default configuration file for Galaxy is now config/galaxy.yml instead of config/galaxy.ini. To minimize the impact of this change on existing Galaxy instances, if a Galaxy has a galaxy.ini file configured, it will continue to use Paste by default unless additional steps are taken by the administrator
Dataset Collection Usability This release has significantly improved the usability of Galaxy dataset collections. Dozens of improvements to collections have been made, some of the key highlights include:
- Data library folders can now be sent to histories as a dataset collection.
- Failed dataset collection elements can now be fixed using job re-running
- Collections now appear with state and progress bars in the history panel and contained datasets are hidden by default
- We added intuitive workflow post job actions for dataset collections
- The web interface now supports collections with arbitrary nesting and size
- More robust nametag discovery and propagation when using collections
Client Architecture The architecture for the client code that powers the Galaxy user interface has been significantly overhauled. The code base has been converted to ES6, Yarn now powers the build and dependency management of the code, Prettier is now used to ensure consistent code formatting, and the VueJS framework has been integrated.
New BAM datatypes Previously Galaxy only supported coordinate sorted BAM files by default (the bam datatype). In addition, this release of Galaxy now supports three new types of BAM:
- qname_sorted.bam, that ensures that the file is queryname sorted (e.g. SO:queryname);
- qname_input_sorted.bam, that can be used to describe the output of aligners which generally keep mate pairs adjacent
- unsorted.bam, that makes no assumptions about the sort order of the file.
Experimental Job Caching Galaxy can now be configured to allow users the option of skipping duplicated jobs if one with identical parameters has been previously executed and simply reuse the previously generated outputs.
See the full release notes for more.
Thanks for using Galaxy!
Galaxy Docker Image 18.01
The Galaxy Docker project has a matching release, for Galaxy 18.01. The release features the 18.01 enhancement and removes nodejs-legacy and npm from the Dockerfile and installs the latest version from ansible-extras.
blend4j 0.2.0
blend4j is a partial reimplemenation of the Python library bioblend for the JVM. bioblend for Python is a library for scripting interactions with Galaxy.
- Update to galaxy-bootstrap 0.7.0.
- Added support for accessing Galaxy Tool Data tables. Thanks to Dan Fornika.
- blend4j now requires Java 1.8+ to run.
galaxy-lib 18.5.3 - 18.5.7
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
Earlier Releases
Other packages released in the prior 4 months.
Planemo 0.48.0
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. These releases included numerous fixes and enhancements.
See GitHub for details.
Pulsar 0.8.1-3
Pulsar updates were released in February. Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
ephemeris 0.8.0
Ephemeris is a small Python library and set of scripts for managing the bootstrapping of Galaxy plugins - tools, index data, and workflows.
- Free software: Academic Free License version 3.0
- Documentation: https://ephemeris.readthedocs.org.
- Code: https://github.com/galaxyproject/ephemeris
blend4php 0.1 beta
The beta version of the blend4php package, a PHP wrapper for the Galaxy API, was released in December. It provides a PHP package for interacting with Galaxy and CloudMan. blend4php currently offers a partial implementation of the Galaxy API and includes support for datasets, data types, folder contents, folders, genomes, group roles, groups, group users, histories, history contents, jobs, libraries, library contents, requests, roles, search, tools, toolshed repositories, users, visualizations and workflows.
The motivation for development of this library is for integration with Tripal, an open-source toolkit for creation of online genomic, genetic and biological databases.
Please see the API documentation page for full information.
And the rest …
Other Galaxy packages that haven’t had a release in the past four months can be found on GitHub.
Other News
- From Yvan Le Bras:
- A demo of the use of Wallace on Galaxy through dockerization, with interaction between Wallace and Galaxy to import from and/or export to your current history, just to enhance the accessibility of Wallace (you don’t need R) plus integration with others tools! Another nice job (did you ever use OpenRefine on Galaxy ?) ) from ValentinChambon !
- From Tomas Klingström:
- We have a new version of Galaksio available. It works well for us but needs to be tested on more different workflows to identify new issues.
- From Stephen Newhouse:
- We have deployed Galaxy on the Genomics England research environment for teaching and then more.
- From Anna Syme:
- The new Genome annotation with Prokka tutorial
- Family Gathering: Galaxy Program Makes Grouping Genes Easier, Technology Networks