On this page
The September 2018 Galactic News is here! This is a summary of what is going on in the Galaxy community. Here’s what’s happening:
- Events
- 2018 SACNAS Workshops
- European Galaxy Days, 19-20 November, Freiburg, Germany
- All upcoming events
- New:
- 80 publications
- Galactic Blog entry: Contributor of the Month: Carrie Ganote
- Public Galaxy Server News: epiGeEC and MetaCentrum
- ToolShed contributions
- Hub and Documentation updates
- CloudBridge v1 has been released!
- Releases
- And other news too
If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events
As always, there is a lot going on in the Galaxy:
2018 SACNAS Pre-conference Workshops
[
 ](/events/2017-sacnas/)
](/events/2017-sacnas/) [
 ](https://software-carpentry.org/)
](https://software-carpentry.org/)[
 ](https://data-carpentry.org/)
](https://data-carpentry.org/)[
 ](https://www.cyverse.org/)
](https://www.cyverse.org/)[
 ](/)
](/)
If you will be anywhere near San Antonio, Texas on October 10 then please consider participating in one of these SACNAS preconference workshops. You don’t need to be registered for the 2018 SACNAS meeting to attend. Everyone is eligible and the workshops are free.
Data Platforms for Life Science Researchers
Registration is free but space is limited. You are strongly encouraged to register soon. Both workshops are held October 10, the day before SACNAS starts.
Plese note: The Galaxy Community Fund is augmenting SACNAS Travel Scholarships awards for the first ~ 28 scholarship recipients that register for these workshops. The additional funds will cover an extra night’s lodging for scholarship recipients, enabling you to arrive the day before the workshops. Funds are limited, and once the funds are allocated, they are gone. (You must have received a SACNAS Travel Scholarship to be eligible.)
European Galaxy Days, 19-20 November, Freiburg, Germany
European Galaxy Days will be held 19 and 20 November 2018 in Freiburg, Germany. The first day will give an overview of the current state of the Galaxy framework with several talks and demonstrations. The second day will focus on developing and extending the Galaxy ecosystem.
The event is free and registration is open. See the event home page for more.
Upcoming events
These and other Galaxy related events are coming up in the next few months:
| Date | Topic/Event | Venue/Location | Contact |
|---|---|---|---|
| September 17th 2018 | 6th Galaxy high-throughput sequencing (HTS) data analysis workshop |  Freiburg Galaxy Team Freiburg Galaxy Team |
|
| September 18th 2018 | Variant detection using Galaxy |  training@qfab.org training@qfab.org |
|
| September 25th 2018 | Gateways 2018 | Katherine Lawrence | |
| October 2nd 2018 | Genome assembly using Galaxy |  training@qfab.org training@qfab.org |
|
| October 4th 2018 | ELIXIR CZ Proteomics Workshop 2018 |  Björn Grüning Björn Grüning |
|
| October 8th 2018 | Workflow4Experimenters 2018 Course |  Gildas Le Corguillé Gildas Le Corguillé |
|
| October 10th 2018 | SACNAS 2018 Pre-Conference Workshops |  Camille Avestruz, Tracy Teal, Blake Joyce, Dave Clements, Joslynn Lee Camille Avestruz, Tracy Teal, Blake Joyce, Dave Clements, Joslynn Lee |
|
| October 15th 2018 | Galaxy @ eResearch Australasia |  Presenters Presenters |
|
| October 15th 2018 | Analyse RNAseq sous Galaxy |  christine.mantecon @upmc.fr christine.mantecon @upmc.fr |
|
| October 16th 2018 | Bioinformatique - Analyse avancée de séquences | Intervenants | |
| October 17th 2018 | Bioinformatics for Translational Medicine using Galaxy: see it, do it, teach it |  Celia van Gelder Celia van Gelder |
|
| October 18th 2018 | Introduction to Galaxy & the Genomics Virtual Laboratory |  Melbourne Bioinformatics Melbourne Bioinformatics |
|
| October 23rd 2018 | Introduction to Galaxy Australia: Differential Gene Expression from Bacterial RNA-seq Data |  |
|
| November 12th 2018 | BioHackathon 2018 Paris | Bérénice Batut, Hervé Ménager | |
| November 13th 2018 | Galaxy for NGS Data Analysis | Weihong Yan | |
| November 14th 2018 | Analyse des données RNA-Seq sous l'environnement Galaxy |  contact@biosciencesco.fr contact@biosciencesco.fr |
|
| November 19th 2018 | RNA-seq analysis in Galaxy | mo@galaxyproject.org | |
| November 19th 2018 | European Galaxy Days | Hans-Rudolf Hotz, Björn Grüning, Jean-François Dufayard | |
| December 6th 2018 | Molecular Dynamics and Analysis using BRIDGE | Organizers | |
| July 1st 2019 | 2019 Galaxy Community Conference (GCC2019) |  Organizers Organizers |
Publications
80 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in August.
Highlighted Publications
Four Galactic and Stellar publications.
- ゲノム情報解析の再現性と再利用性を向上させる情報基盤の設計, thesis by 山中 遼太
- 次世代シーケンサーデータの解析手法 第 11 回 統合データ解析環境 Galaxy, by , 大田 達郎, 寺田 朋子, 清水 謙多郎 , 門田 幸二, Japanese Journal of Lactic Acid Bacteria
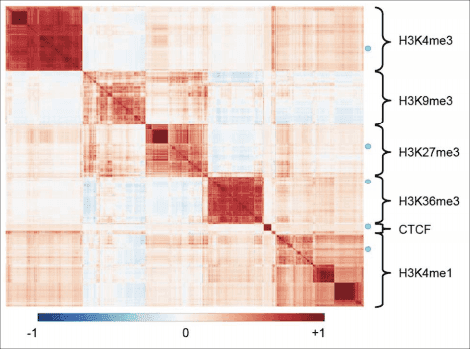
- The epiGenomic Efficient Correlator (epiGeEC) tool allows fast comparison of user datasets with thousands of public epigenomic datasets, by Jonathan Laperle, Simon Hébert-Deschamps, Joanny Raby, David Anderson de Lima Morais, Michel Barrette, David Bujold, Charlotte Bastin, Marc-Antoine Robert, Jean-François Nadeau, Marie Harel, Alexei Nordell-Markovits, Alain Veilleux, Guillaume Bourque, Pierre-Étienne Jacques, Bioinformatics, bty655, doi:10.1093/bioinformatics/bty655
- Bioconda: sustainable and comprehensive software distribution for the life sciences, by Björn Grüning, Ryan Dale, Andreas Sjödin, Brad A. Chapman, Jillian Rowe, Christopher H. Tomkins-Tinch, Renan Valieris, Johannes Köster & The Bioconda Team
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 41 | +Methods | 17 | +Workbench | 13 | +RefPublic | 13 | +UsePublic | |||
| 5 | +UseLocal | 5 | +UseMain | 4 | +Cloud | 4 | +Tools | |||
| 2 | +IsGalaxy | 2 | +Reproducibility | 1 | +HowTo | 1 | +Unknown |
Who’s Hiring

The Galaxy is expanding! Please help it grow.
- Développement d’interopérabilité entre ressources bioinformatique, IFB, France
- Bioinformaticien/biostatisticien biologiste, Phylogene S.A, Bernis, France
- M2 : Diversité bactérienne associée aux microalgues toxiques (metabarcoding), IFREMER Centre Bretagne, Plouzané, France
- Freiburg Galaxy Team has open positions, Freiburg, Germany
- The Blankenberg Lab in the Genomic Medicine Institute at the Cleveland Clinic Lerner Research Institute is hiring postdocs.
- Galaxy Project is hiring software engineers and postdocs at Johns Hopkins, Baltimore, Maryland, United States
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Contributor of the Month: Carrie Ganote
 ](/news/)
](/news/)
There is one new Galactic Blog entry:
- Contributor of the Month: Carrie Ganote, By Björn Grüning
Public Galaxy Server News
There are over 100 publicly accessible Galaxy servers and several Galaxy services. One of each were added in the past month.
epiGeEC
epiGeEC compares user-provided epigenomic datasets with thousands of public datasets in a few minutes. It can also directly compare user’s datasets. This server is provided by the Genetics and genomics Analysis Platform (GenAP) project, thereby leveraging Compute Canada’s advanced research computing infrastructure. A tutorial is available and anonymous access and account creation are both supported.”
See the epiGeEC directory entry for more.
MetaCentrum - Czechia
MetaCentrum supports Galaxy instances on the Czech National Grid Infrastructure (NGI), that are available to employees and students of research and academic organizations of Czechia and for scientific purposes only. Use requires registering for an account.
See the Metacentrum directory entry for more.
Public Servers in Publications
We tag papers that use, mention, implement or extend public Galaxy Servers. Here are the counts for the past month’s publications:
Hub and Doc Updates
- The Galaxy publication library is now integrated into the Pan-Galactic search page, thanks to Dannon Baker. Search for something and click on the pub library tab.
- Data Manager XML syntax description updated by Peter van Heusden
- Hub Editing Guidelines updated by Mo Heydarian
- There was a massive Hub infrastructure update by Dannon Baker, but it’s one of those you won’t even notice.
Tools
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in August 2018.
Releases
New additions to the Galaxy Community.
CloudBridge
CloudBridge v1 has been released!
- One interface to major Infrastructure-as-a-Service (IaaS) cloud providers
- CloudBridge v1 supports Amazon Web Service (AWS), Microsoft Azure, and OpenStack clouds
- Thanks to help from the AnswerALS project and Microsoft, Azure now has complete implementation of all CloudBridge capabilities
For other features and changes contained in this release, please take a look at the CHANGELOG.
BioMAJ2Galaxy
BioMAJ2Galaxy does the following actions on a Galaxy server:
- Add new items to data libraries
- Remove items from data libraries
- Add new items to tool data tables using data managers
- Remove items from tool data tables
These scripts are primarily designed to be used as BioMAJ post processes, but they can probably used directly from the command line if you need to.
Galaxy Docker Image 18.05
The Galaxy Docker project has a matching release, for Galaxy 18.05. The release features the 18.05 enhancements.
And the rest …
Other Galaxy packages that haven’t had a release in the past four months can be found on GitHub.
Other News
- CellOrganizer for Galaxy is a set of tools that enables users to train generative models of the cell from microscopy images, analyze trained generative models, and synthesize instances using CellOrganizer, a tool for learning and managing generative models of cell organization directly from images.