October 2019 Galactic News
Events! Pubs! Platforms! Jobs! Doc! Tutorials! And some other news too.
On this page
The October 2019 Galactic News includes Galactic news under this or any other sun:
- 20 Upcoming Events
- 143 new publications
- Galaxy Platform News
- 6 Openings
- Training material and doc updates
- Other news
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events

GalaxyAfrica 2019 will be 14-15 November, in collaboration with the ASBCB/ISCB Africa conference in Kumasi, Ghana. We will offer Galaxy training for researchers, and training for systems administrators who need to maintain a local installation. This year we are calling for abstracts for three oral presentations that demonstrate the use of Galaxy in your research projects.
Early registration ends 20 October.

This year’s Galaxy Netherlands Face2Face meeting will take place at:
- Time: October 3, 13:30-16:00, with drinks afterwards
- Location: GROW room, DTL offices, JIM 6th floor, Jaarbeursplein 6, 3521 AL Utrecht
Interested? Please register for the meeting.
Brussels, Belgium. Monday 3th to Friday February 7th 2020.
This one-week course (entirely in English), Learn to use the W4M infrastructure and analyze your own LC-MS, GC-MS, or NMR data. Morning sessions will be dedicated to methodology and tools. Afternoon sessions will be devoted to tutoring.
Preregister by 3 November.

There are
- 20 upcoming events
- on 5 continents
- in Italy, Germany, the Netherlands, Greece, Estonia, Belgium, US, France, Taiwan, Ghana, Australia, and UK.
- on RNA-Seq, bisulfite sequencing, reproducibility, CWL, cancer, single cell, metabarcoding, and metabolomics!

Galaxy, differential gene expression analysis, bisulfite sequencing analysis, and Nanopore data exploration and usage. Three days in Freiburg starting 9 October.

Reproducible Genomic Data Analysis with the Galaxy Workbench, 9:30 - 11:20, Friday, October 11.

Reproducible and Transparent Genomic Analysis with Galaxy, Tuesday, October 15, 1:30 - 3:00. Register now, as space is limited.

Savoir utiliser l’environnement Galaxy, à partir du 29 octobre.
There are several registration deadlines this month:
- Galaxy Africa & ISCB Africa, Kumasi, Ghana. Early Registration: October 20
- W4E 2020, Brussels, Belgium. Pre-registration: 3rd November
Publications
143 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month. There were 9 Galactic and Stellar publications added, and four of those are open access:
Andrio, Pau, Adam Hospital, et al. 2019. Scientific Data 6 (1): 1–8. doi: 10.1038/s41597-019-0177-4.
Andersen, Thea Os. 2019. Thesis, Norwegian University of Life Sciences.
Ellrott, Kyle, Alex Buchanan, et al. Genome Biology 20 (1): 195. doi: 10.1186/s13059-019-1794-0. 2019.
Daniel Blankenberg, Martin Cech. 2019. In Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics (BCB ‘19). ACM, New York, NY, USA, 559-559. DOI: 10.1145/3307339.3343178
Publications are tagged with how they use, extend or reference Galaxy. The past month’s pubs were tagged as:
104 : Methods 46 : UsePublic 15 : Workbench 12 : UseLocal 11 : UseMain 8 : Tools 7 : RefPublic 4 : IsGalaxy 2 : Cloud 2 : Other 2 : Project 2 : Reproducibility 1 : HowTo
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. There were are many new platforms this month:

The Inter-Domain Ecological Network Analysis Pipeline (IDENAP) server models, analyzes and manages the Inter-Domain Ecological Networks (IDENs) from metagenomic datasets, e.g. GeoChip and Sequencing, and synchronous plant surveys.

The ELIXIR Belgium Galaxy server has been launched. This server focuses on serving researchers in Belgium, provides a wealth of tools, and reference data for plant species from the PLAZA data warehouse.

Lebanese University has launched the first public Galaxy server in the Middle East. The server features the SpCLUST tool set for fast and reliable clustering of potentially divergent biological sequences. It also supports the CD-Hit and DNAClust tool sets.

The NG-Tax server provides a semantic framework for high-throughput marker gene (amplicon) analysis. The server requires that you create a login, but anyone can create a login. Supports both NG-Tax versions 1 & 2.

Galaxy for Ecology is a web platform to get, process, analyze and visualize ecological data, including species occurrence data, protocoled data from Vigie-Nature, and climatic data from Worldclim.

Platforms that were referenced at least twice in the past month’s publications:
23 : Huttenhower 5 : Workflow4Metabolomics 4 : ARGs-OAP 4 : UseGalaxy.eu 3 : Cistrome 3 : Pasteur 2 : AmrPlusPlus 2 : RepeatExplorer 2 : UseGalaxy.org.au
Who’s Hiring
The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back! Have a Galaxy-related opening? Send it to outreach @ galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Full-Time, Rixensart, Belgium.
Develop core bioinformatics pipelines using a variety of environments (incl. Linux, Galaxy, DNA-Nexus) for base-calling, demultiplexing, data quality control, sequence alignment, variant calling, quantifying gene expression…

The Plant Computational Genomics lab in the Department of Ecology and Evolutionary Biology at the University of Connecticut seeks motivated MS and PhD students to join the lab in the Summer/Fall 2020.

AbSci, Vancouver, Washington, United States

Interested in an exciting new Galaxy-focused opportunity with core team members? Contact recruiting@galactic-core.com today.

At the VIB-UGent Center for Plant Systems Biology, with ELIXIR Belgium.

The The European Galaxy Team has open positions in Freiburg, Germany.

Doc, Hub, and Training Updates
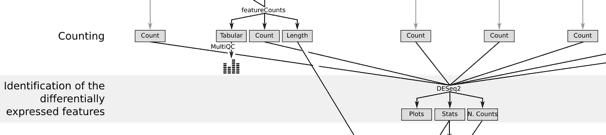
The [Reference-based RNA-Seq data analysis](https://training.galaxyproject.org/topics/transcriptomics/tutorials/ref-based/tutorial.html) has been expanded by [Bérénice Batut](https://training.galaxyproject.org/training-material/hall-of-fame#bebatut), [Anika Erxleben](https://training.galaxyproject.org/training-material/hall-of-fame#erxleben), and [Nicola Soranzo](https://training.galaxyproject.org/training-material/hall-of-fame#nsoranzo) to provide more details about DGE, GO and KEGG analysis, and visualization.

By [Katarzyna Murat](https://training.galaxyproject.org/training-material/hall-of-fame#kpbioteam) and [Krzysztof Poterlowicz](https://training.galaxyproject.org/training-material/hall-of-fame#kpoterlowicz)
This tutorial identifies differentially methylated regions and positions associated with treatment resistant melanomas.
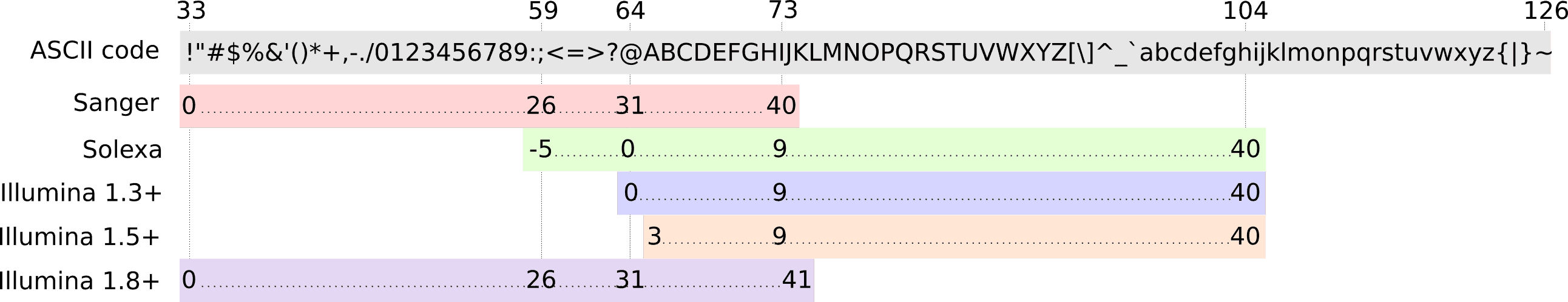
The [Quality Control tutorial](https://training.galaxyproject.org/topics/sequence-analysis/tutorials/quality-control/tutorial.html) has been updated by [Bérénice Batut](https://training.galaxyproject.org/training-material/hall-of-fame#bebatut), [Saskia Hiltemann](https://shiltemann.github.io/), [Anup Kumar](https://github.com/anuprulez), Matthias (which one, we ask?) and [Anika Erxleben](https://github.com/erxleben). It provides more details about FastQC reports and clarifies the trimming approach and paired-end explanation.

[Galaxy Admin](https://training.galaxyproject.org/training-material/topics/admin/) & [Dev](https://training.galaxyproject.org/training-material/topics/dev/) topics now include Matrix chat rooms embedded into the tutorials. Ask your questions or just start a discussion about the training topics with fellow learners and tutorial authors.

By [Valentin Loux](https://training.galaxyproject.org/training-material/hall-of-fame#vloux), [Florence Combes](https://training.galaxyproject.org/training-material/hall-of-fame#combesf), [David Christiany](https://training.galaxyproject.org/training-material/hall-of-fame#davidchristiany), [Yves Vandenbrouck](https://training.galaxyproject.org/training-material/hall-of-fame#yvandenb).
Annotate a protein list identified by LC-MS/MS experiments using the [ProteoRE Galaxy instance](http://www.proteore.org/).

By [Mehmet Tekman](https://training.galaxyproject.org/training-material/hall-of-fame#mtekman), [Hans-Rudolf Hotz](https://training.galaxyproject.org/training-material/hall-of-fame#hrhotz), and [Daniel Blankenberg](https://training.galaxyproject.org/training-material/hall-of-fame#blankenberg)
Map and quantify scRNA-seq FASTQ data via RNA STARsolo, and reproduce a Cell Ranger workflow using the DropletUtils suite. Explore the use of barcode rankings to determine better filtering thresholds to generate a high quality count matrix.

By [Lucille Delisle](https://training.galaxyproject.org/training-material/hall-of-fame#lldelisle), [Maria Doyle](https://training.galaxyproject.org/training-material/hall-of-fame#mblue9), [Florian Heyl](https://training.galaxyproject.org/training-material/hall-of-fame#heylf)
Learn ATAC-Seq analysis with data from the study of Buenrostro et al. 2013, the first paper on the ATAC-Seq method. Compare predicted open chromatin regions to known binding sites.
Other News
Galaxy is participating in HacktoberFest, the annual drive for global contribution in the growth of open source software has begun. Sign up and make a pull request for documentation, new tools, or contribute to any of the Galaxy GitHub repositories to earn free GitHub and digitalocean swag.

The GenomeSpace project ends on November 15, 2019 due to expiration of its NHGRI funding and we will be shutting down the GenomeSpace servers on that date. Galaxy was a GenomeSpace-enabled tool from the beginning of GenomeSpace and worked closely to maintain that connection over the years.

The latest release of InterMine adds configuration free data import/export with Galaxy. An InterMine tutorial in the Galaxy Training Network is coming…
See the full release announcement and release notes for more on this release.


