Galactic December News!
On this page
Welcome to the December 2015 Galactic News, a summary of what is going on in the Galaxy community.
If you have anything to include in the next News, please send it to [Galaxy Outreach](mailto:outreach AT galaxyproject DOT org).
New Papers
81 new papers referencing, using, extending, and implementing Galaxy were added to the Galaxy CiteULike Group in November. Highlights include:
-
Genomics Virtual Laboratory: A Practical Bioinformatics Workbench for the Cloud by Enis Afgan, Clare Sloggett, Nuwan Goonasekera et al., PloS One, Vol. 10, No. 10. (2015)
-
Cost-Aware Cloud Provisioning by Ryan Chard, Kyle Chard, Kris Bubendorfer, Lukasz Lacinski, Ravi Madduri, Ian Foster, In e-Science (e-Science), 2015 IEEE 11th International Conference on (August 2015), pp. 136-144, doi:10.1109/escience.2015.67
-
DNA data bank of Japan (DDBJ) progress report by Jun Mashima, Yuichi Kodama, Takehide Kosuge, et al. Nucleic Acids Research (17 November 2015), gkv1105, doi:10.1093/nar/gkv1105
-
antaRNA – Multi-objective inverse folding of pseudoknot RNA using ant-colony optimization by Robert Kleinkauf, Torsten Houwaart, Rolf Backofen, Martin Mann, BMC Bioinformatics, Vol. 16, No. 1. (18 November 2015), 389, doi:10.1186/s12859-015-0815-6
The new papers were tagged with:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 4 | Cloud | 1 | Project | 5 | Tools | 14 | UsePublic | |||
| 1 | HowTo | 6 | RefPublic | - | UseCloud | - | Visualization | |||
| 7 | IsGalaxy | 2 | Reproducibility | 3 | UseLocal | 23 | Workbench | |||
| 37 | Methods | 1 | Shared | 11 | UseMain |
Events
Spanish Galaxy Community Meetup: 18 December, Barcelona
There will be a session at the end of the III Bioinformatics and Computational Biology Symposium being held in Barcelona on 18 December:
where users and developers can, not only share information and code, but also plan how to coordinate for future Galaxy workshops and other activities. Chairs of this session are Toni Espinosa (UAB) and Gonzalo Vera (CRAG).
See the announcement PDF and the announcement email for further details.
Galaxy Day 2015 Presentations
Galaxy Day 2015 was held in Paris on 19 November. It was preceded by a Galaxy Hackathon. Both events were well attended and the events were yet another successful gathering of the French Galaxy community. Presentations from Galaxy Day and a summary of the Hackathon are now available online (click on “Archives Galaxy”).
GCC2016 Registration will open this month
GCC2016 registration will open this month. The full training schedule will also be posted at that time. We’ll announce this on all Galaxy channels - you won’t miss it.
Upcoming Events
There are many upcoming events in the next few months. See the Galaxy Events Google Calendar for details on other events of interest to the community.
| |
Designates a training event offered by GTN member(s) |
Who’s Hiring
The Galaxy is expanding! Please help it grow.
- intégration d’outils phylogénétiques dans Galaxy, ANSES Ploufragan, Unité de Génétique Virale et Biosécurité, France
- Computational Biologist / Bioinformatician, The Sainsbury Laboratory, Norwich, UK.
- Software Engineer, Oregon Health Sciences University, Portland, Oregon, United States
- Software Developer / Bioinformatician, European Molecular Biology Laboratory (EMBL), Heidelberg, Germany Closes 13 December 2015.
- The Galaxy Project is hiring software engineers and post-docs
Got a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
New Public Galaxy Servers
Two new publicly accessible Galaxy servers were added in November, bringing the total precipitously close to 80.
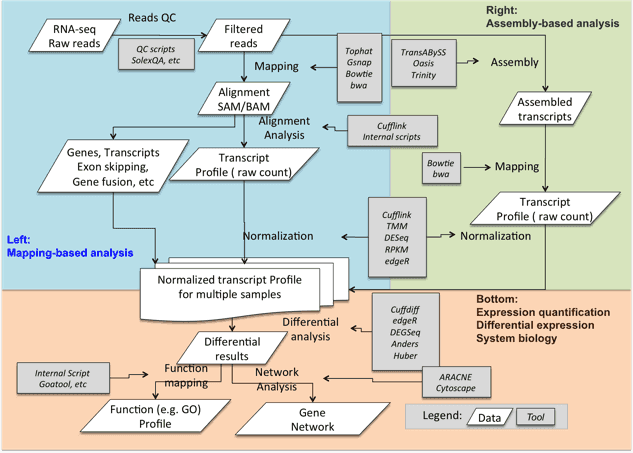
Galaxy Integrated Omics (GIO)
- Link:
- Domain/Purpose:
- “Proteomics Informed by Transcriptomics (PIT) methodology described in Evans et al. 2012, and selection of surrogate peptides for targeted proteomics.”
- Comments:
- “Galaxy-based Integrated Omics (GIO) is a curated collection of new and pre-existing open source tools brought together for proteomics applications.”
- See Galaxy Integrated Omics: Web-based standards-compliant workflows for proteomics informed by transcriptomics, Jun Fan, Shyamasree Saha, Gary Barker, Kate J. Heesom, Fawaz Ghali, Andrew R. Jones, David A. Matthews and Conrad Bessant, Molecular & Cellular Proteomics, 14, 3087-3093.
- User Support:
- Tutorial 1:Getting started with GIO
- Tutorial 2:Getting started with GIO workflows
- Tutorial 3:Proteomics Informed by Transcriptomics (PIT)
- Tutorial 4:Targeted proteomics
- For draft documentation please visit the GIO repository.
- Quotas:
- Sponsor(s):
- “This project is led by Conrad Bessant at Queen Mary and David Matthews at Bristol, with additional contributions from the groups of Andy Jones at Liverpool and Simon Hubbard at Manchester. GIO development was supported by BBSRC TRDF2 grants BB/L018438/1 (Proteomics Goes Viral), BB/K016075/1 (Galaxy Workflows for Proteomics Informed by Transcriptomics) and BB/K004123/1 (Integrating Genomes and Proteomes on the Cloud). Lead developer is Jun Fan.”
RNA-Seq Portal
-
Links:
-
Domain/Purpose:
- Analyzing RNA-seq Data for Agricultural Animal Species.
-
Comments:
- From the portal landing page:
The goal of this project is to develop a web portal with integrated tools for RNA-seq based gene expression analysis for agricultural animals.
- improve genome annotation of agricultural animal species, including (but not limiting to) cattle, pig, chicken, turkey, horse, sheep, and goat as well as catfish;
- develop and integrate needed bioinformatics tools and pipelines, visualization interfaces, and statistical methods;
- build a web portal that enable RNA-seq based transcriptomics analysis in aforementioned animal species.
- From the portal landing page:
The goal of this project is to develop a web portal with integrated tools for RNA-seq based gene expression analysis for agricultural animals.
-
User Support:
- Example usage (see center panel)
- Web support
-
Quotas:
-
Sponsor(s):
- The project is developed by Weizhong Li’s lab at J. Craig Venter Institute with support from NIFA (award #2013-67015-21428).
Releases
Planemo 0.19.0 - 0.21.1
Planemo saw a regular Oort cloud of activity in November with 3 releases.
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. Planemo 0.21.1 is the most recent release. See the release history.
Other Releases
July 2015 Galaxy Release (v 15.07)
Galaxy Release v 15.07 includes enhancements to Interactive Environments and the Workflow Editor, an update of the Policies for Committers and Pull Requests and much more.
BioBlend 0.7.0
BioBlend version 0.7.0 was released at the beginning of November. BioBlend is a python library for interacting with CloudMan and the Galaxy API. CloudMan offers an easy way to get a personal and completely functional instance of Galaxy in the cloud in just a few minutes, without any manual configuration.) From the release CHANGELOG.
Mid 2015 Galaxy CloudMan Release
Security Advisories
Three security vulnerabilities were announced and patched in August. The Galaxy Committers have released a number of fixes which have been rolled in to affected versions of Galaxy dating back to the 14.10 release.
Pulsar Pulsar 0.5.0 was released in May. Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
blend4j v0.1.2 blend4j v0.1.2 was released in December 2014. blend4j is a JVM partial reimplemenation of the Python library bioblend for interacting with Galaxy, CloudMan, and BioCloudCentral.
Galaxy Community Hubs
 |
 |
 |
| Share your training resources and experience now | Share your experience now |
ToolShed Contributions
See list of tools contributed in October and November.
Other News
- From Guy Reeves: ‘Hidden’ Galaxy features which are really useful. For Scratchbook see this video.
- Ever wondered who writes Galaxy? These people do.
- See the executive summary of free open source software the Galaxy community provides. (Big thanks to Helena Rasche.)
- From Nate Coraor An Ansible role for installing Galaxy Interactive Environments
- Expanding the Galaxy Platform at the Minnesota Supercomputing Institute.
- From Björn Grüning: The Galaxy Docker Image has gained TravisCI testing. BioBlend will test all functionality and tool installation.
- Updated documentation: