It’s a new year in the Galaxy!
On this page
Welcome to the January 2016 Galactic News, a summary of what is going on in the Galaxy community.
If you have anything to include in the next News, please send it to [Galaxy Outreach](mailto:outreach AT galaxyproject DOT org).
New Papers
51 new papers referencing, using, extending, and implementing Galaxy were added to the Galaxy CiteULike Group in December. Highlights include:
-
Sequence database versioning for command line and Galaxy bioinformatics servers by Damion Dooley, Aaron Petkau, Gary Van Domselaar, William Hsiao, Bioinformatics (12 December 2015), btv724, doi:10.1093/bioinformatics/btv724
-
Dintor: functional annotation of genomic and proteomic data by Christian X. Weichenberger, Hagen Blankenburg, Antonia Palermo, et al. BMC Genomics, Vol. 16, No. 1. (21 December 2015), doi:10.1186/s12864-015-2279-5
-
Globus Nexus: A Platform-as-a-Service provider of research identity, profile, and group management by Kyle Chard, Mattias Lidman, Brendan McCollam, et al. Future Generation Computer Systems, Vol. 56 (March 2016), pp. 571-583, doi:10.1016/j.future.2015.09.006
-
Enhanced reproducibility of SADI web service workflows with Galaxy and Docker by Mikel Egaña E. Aranguren, Mark D. Wilkinson GigaScience, Vol. 4, No. 1. (3 December 2015), doi:10.1186/s13742-015-0092-3
-
FuMa: reporting overlap in RNA-seq detected fusion genes by Youri Hoogstrate, René Böttcher, Saskia Hiltemann, Peter van der Spek, Guido Jenster, Andrew P. Stubbs, Bioinformatics (10 December 2015), btv721, doi:10.1093/bioinformatics/btv721
-
ADMIRE: analysis and visualization of differential methylation in genomic regions using the Infinium HumanMethylation450 Assay by Jens Preussner, Julia Bayer, Carsten Kuenne, Mario Looso Epigenetics & chromatin, Vol. 8, doi:10.1186/s13072-015-0045-1
The new papers were tagged with:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | Cloud | - | Project | 8 | Tools | 9 | UsePublic | |||
| 2 | HowTo | 7 | RefPublic | - | UseCloud | - | Visualization | |||
| 6 | IsGalaxy | - | Reproducibility | 2 | UseLocal | 12 | Workbench | |||
| 23 | Methods | 1 | Shared | 8 | UseMain |
Events
GCC2016 Website is online
The 2016 Galaxy Community Conference website is now online and includes registration information and the full training schedule.
GCC2016 will be held June 25-29 at Indiana University in Bloomington, Indiana, United States. This will be the 7th annual gathering of the Galaxy community, and we are expecting over 200 participants again this year.
Registration information describes the different registration packages, rates, and deadlines. Early registration rates start at less than $40 per day for students and post-docs.
The training schedule includes session titles, descriptions, prerequisites, and instructors. Many thanks to everyone who nominated and voted on topics, and especially to the many instructors who are contributing their time and effort to make GCC2016 a success.
Thanks also go to the Indiana University Pervasive Technology Institute (PTI), who are supporting GCC2016 as a Platinum Sponsor. PTI is home to the National Center for Genome Analysis Support (for which NSF funding was recently renewed), and the Galaxy Trinity instance, and is a long time supporter of the Galaxy Community. PTI pairs fundamental academic computational research with the widely known strengths of Indiana University through innovations and service delivery in networking and high performance computing. See the PTI website for more information.
Seeking Sponsors
We continue to seek other sponsors as well and offer a wide range of sponsorship plans. If your organization is interested in having a presence at GCC2016, please contact the GCC2016 Exec for more information.
Registration will open this month
Yeah, we know, we said this last month. But last month “this month” actually meant next month which is now this month, due to, ummm, failure to compensate for the 2015 leap second. Yeah.
Anyway, watch Galaxy channels for registration opening this month, really.
Metagenomics Tools and Workflows Codefest Reports
The Intergalactic Utilities Commission held it’s second codefest with the aim to improve metagenomic research in Galaxy.
The IUC was available for two days in a Hangout to answers questions and to work together with the Galaxy community to enhance our tools and the overall Galaxy ecosystem. 22 scientists from all over the world participated in this and made it a fantastic experience and a success story, which the IUC would like to repeat more often.
There are two event reports, See the one by Björn Grüning, and one by Yvan Le Bras.
Upcoming Events
There are many upcoming events in the next few months. See the Galaxy Events Google Calendar for details on other events of interest to the community.
| |
Designates a training event offered by GTN member(s) |
Who’s Hiring
The Galaxy is expanding! Please help it grow.
- Computational Biology post-doc, IMI-eTRIKS consortium, CNRS-EISBM, Lyon France. Contribute “to the development of eTRIKS Galaxy tools and workflows for disease stratification and biomarker discovery from single and multiple omics datasets.”
- Post-doc in Functional Genomics Of Obesity-Related Diseases, Inserm, Lille, France
- Bioinformatician Post-doc at SB Roscoff working on parasitic dinoflagellate genomics
- Software Engineer, Oregon Health Sciences University, Portland, Oregon, United States
- Software Developer / Bioinformatician, European Molecular Biology Laboratory (EMBL), Heidelberg, Germany Extended until 10 January 2016.
- The Galaxy Project is hiring software engineers and post-docs
Got a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
New Public Galaxy Servers
Two new publicly accessible Galaxy servers were added in December. And, while not a new server, the server formerly known as MIRPIPE is now the greatly expanded MPI-HLR Bioinformatics Server.
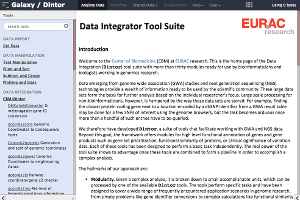
Dintor
-
Link:
-
Domain/Purpose:
- A “suite of tools that facilitate working with GWA and NGS data” and “offers modules for high level functional annotation of genes and gene products such as gene set prioritization, functional similarity of proteins, or clinical significance of variation data. Each of these tools has been designed to perform a basic task independently.”
-
Comments:
- Includes “more than thirty modules ready for use by bioinformaticians and biologists working in genomics research.”
- “The Galaxy server has been set up to facilitate access to our
Dintortools by biologists with little background in bioinformatics. A second, expert mode of invocation is given by command line access to the tool suite, which can be downloaded” - Admin and deployment documentation is available.
- Dintor: functional annotation of genomic and proteomic data, Christian X. Weichenberger, Hagen Blankenburg, Antonia Palermo, Yuri D’Elia, Eva König, Erik Bernstein and Francisco S. Domingues, BMC Genomics 201516:1081 DOI: 10.1186/s12864-015-2279-5
-
User Support:
- Tutorials
- In addition to help on each of the Galaxy Tool pages, there is additional help for each Dintor tool here.
-
Quotas:
- Anyone can create a login. Anonymous use is also supported.
-
Sponsor(s):
Erasmus MC
-
Link:
-
Domain/Purpose:
- General purpose genomics analysis, featuring many standard tools plus many additional tools.
-
Comments:
- See FuMa: reporting overlap in RNA-seq detected fusion genes, by Youri Hoogstrate, René Böttcher, Saskia Hiltemann, Peter van der Spek, Guido Jenster and Andrew P Stubbs, Bioinformatics (2015) doi: 10.1093/bioinformatics/btv721
-
User Support:
-
Quotas:
- “This Galaxy instance may be used without registration, but guests’ histories are deleted nightly. To request an account, please contact the Bioinformatics department.”
-
Sponsor(s):
Releases
Galaxy 15.10
The October 2015 Galaxy Release (v 15.10) was released on November 30.* Highlights include
Reports Application The reports web application has been greatly enhanced - including improved styling, new landing page, more reports, pagination, and sparklines. Huge thanks to Daniel Bouchard (@Airistotal) for these enhancements.
Upload The Galaxy upload widget now features support for composite datatypes and improved styling.
Data Libraries Improved API and UI for data libraries - including support for library folder management and search.
Deprecations:
Mercurial is deprecated.
Old Data Libraries UI is deprecated
graphview visualization is deprecated
Direct access to Tool Shed repositories through the Mercurial API is deprecated
and many smaller updates and bug fixes.
See the full release notes for details.
Galaxy Docker Image 15.10
And, thanks to Björn Grüning, there is also now a Docker image for Galaxy 15.10 as well.
* Hey, we’re the Galaxy Project - we can warp time, dang it!
galaxy-lib 16.1.0 - 16.1.7
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
It is also quite popular and has been downloaded from PyPi almost 4000 times since it was created on December 20.
CloudMan 15.12
CloudMan is a cloud manager that orchestrates all of the steps required to provision a complete compute cluster environment on a cloud infrastructure; subsequently, it allows one to manage the cluster, all through a web browser.
This minor CloudMan 15.12 release includes:
- Update Galaxy to the Galaxy 15.10 release (see above for all the juicy details with the improvements that brings along)
- Preload the GIE IPython Docker container onto the image for faster startup: less than 30 seconds to a fully functional IPython notebook vs. several minutes before!
- For those of you that ran into the problem of the size of your user data overfilling after having added and removed a bunch of worker instances - no worries, that has been fixed now and resource tags are no longer stored in the cluster config to prevent persistent_data.yaml from becoming too big.
- The file system archive extraction has been made more resilient and synchronous.
- For those that are building custom images, a more direct method for locating the Nginx configuration directory has been added to help with different versions of the operating system (thanks to Matthew Ralston)
- A number of dependent library versions have been updated to their latest versions.
Pulsar 0.6.0 - 0.6.1
Pulsar 0.6 was released in December. Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
The 0.6.x release includes these changes:
- Pulsar now depends on the new galaxy-lib Python package instead of manually synchronizing Python files across Pulsar and Galaxy.
- Numerous build and testing improvements.
- Fixed a documentation bug in the code (thanks to @erasche). e8814ae
- Remove galaxy.eggs stuff from Pulsar client (thanks to @natefoo). 00197f2
- Add new logo to README (thanks to @martenson). abbba40
- Implement an optional awknowledgement system on top of the message queue system (thanks to @natefoo). Pull Request 82 431088c
- Documentation fixes thanks to @remimarenco. Pull Request 78, Pull Request 80
- Fix project script bug introduced this cycle (thanks to @nsoranzo). 140a069
- Fix config.py on Windows (thanks to @ssorgatem). Pull Request 84
- Add a job manager for XSEDE jobs (thanks to @natefoo). 1017bc5
- Fix pip dependency installation (thanks to @afgane) Pull Request 73
Other Releases
Planemo 0.21.1
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. Planemo 0.21.1 is the most recent release. See the release history.
BioBlend 0.7.0
BioBlend version 0.7.0 was released at the beginning of November. BioBlend is a python library for interacting with CloudMan and the Galaxy API. CloudMan offers an easy way to get a personal and completely functional instance of Galaxy in the cloud in just a few minutes, without any manual configuration.) From the release CHANGELOG.
blend4j v0.1.2 blend4j v0.1.2 was released in December 2014. blend4j is a JVM partial reimplemenation of the Python library bioblend for interacting with Galaxy, CloudMan, and BioCloudCentral.
Galaxy Community Hubs
 |
 |
 |
| Share your training resources and experience now | Share your experience now |
ToolShed Contributions
See list of tools contributed in December.
Other News
- Help the MetaboHub / Workflow4metabolomics and the University of Birmingham / Galaxy-M teams! Please fill out the Galaxy Workflows for Metabolomics Questionnaire
- From GigaScience:
- We’ve been busy in 2015! Read our end of year blog and we have more exciting things lined up for 2016!