On this page
The November 2019 Galactic News includes Galactic news under this or any other sun:
- 19 Upcoming Events
- 208 new publications
- 2 new blog posts
- Galaxy Platform News
- Training material and doc updates
- Openings with 5 organizations
- Releases
- Other news
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events

GalaxyAfrica 2019 will be 14-15 November, in collaboration with the ASBCB/ISCB Africa conference in Kumasi, Ghana. We will offer Galaxy training for researchers, and training for systems administrators who need to maintain a local installation. This year we are calling for abstracts for three oral presentations that demonstrate the use of Galaxy in your research projects.

21st of November, Online
A dedicated day of collaborative work on the training content and calls with the Galaxy training community. Anyone who would like to contribute, learn how to contribute to the Galaxy Training Material or just catch up with the community is very welcome to join.

Galaxy will be at Plant and Animal Genome XXVII (PAG 2020), in San Diego, California, United States, January 11-15. This includes a hands-on Galaxy Workshop (highlighting the new Excellence in Breeding platform and many talks and posters featuring Galaxy.
And there will be a GMOD Codefest before PAG and an NCBI Codeathon after PAG.

Galaxy will be at 2020 ABRF meeting, in Palm Springs, California, United States, February 29 through March 3. This includes a full day hands-on Galaxy Workshop about using Galaxy with your single cell and microbiome data. ABRF bills itself as “the annual conference for technology-enabled multidisciplinary research.”

Save these dates! The next Galaxy Admin Training will be offered 2-6 March at the Barcelona Supercomputing Center.
This week-long hands-on training will feature what you need to know to set up your own production quality Galaxy server. Registration will open later this month.

There are
- 16 upcoming events
- on 4 continents, plus online
- in US, Ghana, France, Australia, UK, Belgium, and Spain.
Publications
208 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month. There were eight Galactic and Stellar publications added, and six of those are open access:
Schmidt, L., Werner, S., Kemmer, T., Niebler, S., Kristen, M., Ayadi, L., … Helm, M. (2019). Frontiers in Genetics, 10. doi: 10.3389/fgene.2019.00876
Vekris, A., Pilalis, E., Chatziioannou, A., & Petry, K. G. (2019). Frontiers in Physiology, 10. doi: 10.3389/fphys.2019.01160
Ison, J., Ménager, H., Brancotte, B., Jaaniso, E., Salumets, A., Raček, T., … Ienasescu, H.-I. (2019). Briefings in Bioinformatics. doi: 10.1093/bib/bbz075
Mei, H., Arbeithuber, B., Cremona, M. A., DeGiorgio, M., & Nekrutenko, A. (2019). Genome Biology and Evolution. doi: 10.1093/gbe/evz197
McDermott, B. T., Peffers, M. J., McDonagh, B., & Tew, S. R. (2019). Journal of Biological Chemistry, 294(35), 13027–13039. doi: 10.1074/jbc.RA118.006865
Silva, B. S. M. L., Heringer, P., Dias, G. B., Svartman, M., & Kuhn, G. C. S. (2019). BioRxiv, 781146. doi: 10.1101/781146
Publications are tagged with how they use, extend or reference Galaxy. The past month’s pubs were tagged as:
142 : Methods 75 : UsePublic 25 : UseMain 24 : Workbench 15 : RefPublic 10 : UseLocal 7 : Reproducibility 6 : Tools 5 : Other 5 : Shared 3 : IsGalaxy 2 : Cloud 1 : HowTo
Galactic Blog Activity
Two new entries in the past month, both related to teaching how to do stuff.
A howto guide on using the new Table Compute tool to: drop or duplicate given rows or columns from a tabular dataset, filter rows or columns on their properties, replace or transform table element, compute tables of summary statistics across rows or columns,…
Agricultural Sciences collaboration between Swedish University of Agricultural sciences researchers and the Agricultural Research Council in South Africa.
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. There are many new platforms this month:

A general purpose Galaxy server optimized for microbiological genomics applications of interest for public health. Support analysis of Illumina and IonTorrent data. A set of YouTube videos will be released in the beginning of 2020.

Hosts tools for the GOBii project, HTPG project and EiB platform, including genomic selection, marker selection, GWAS, imputation, file conversion, and cluster analysis. CGIAR supports EiB to help modernize breeding programs targeting the developing world for greater impact on food and nutrition security, climate change adaptation and development.

A general purpose genomics server with an emphasis on metagenomics, including FROGS and several Migale developed tools. GeDI maps shotgun metagenomes on reference genomes for estimating microbial ecosystems; and MetaFoldScan scans metagenomes and identifies fold hits associated to a structurally characterized target protein. See the Migale newsletter for more.
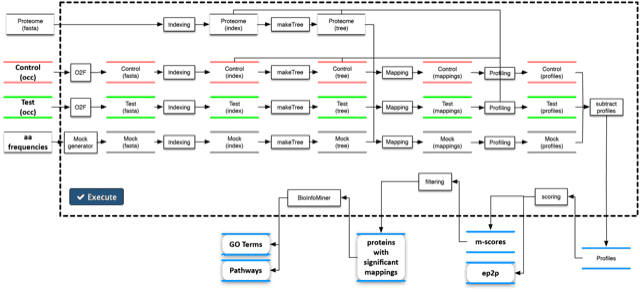
Mapping of massive peptide repertoires on whole proteomes and delivering streamlined, systems-level biological interpretation. PepSimili provides a systems-level interpretation of the mechanisms impacted by the cumulative effect of multiple mimicking peptides on protein networks. It shortlists and ranks candidate target proteins derived from Phage Display experiments.
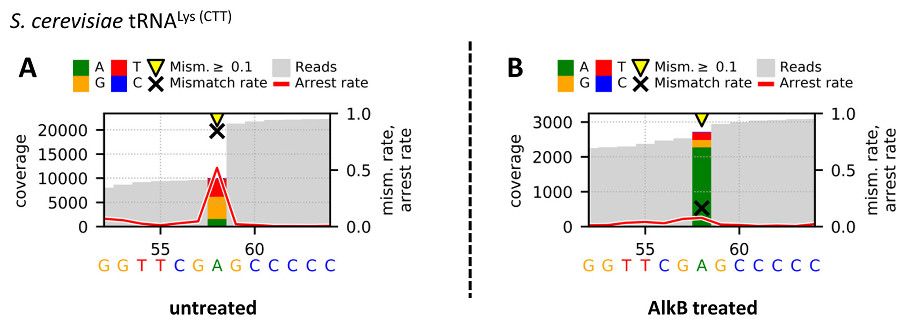
A Galaxy Docker image for RNA analysis and modification calling, from the Helm Group of the Institute of Pharmacy and Biochemistry at Johannes Gutenberg-University.

- UseGalaxy.eu Tool Updates for 5th, 12th, 17th, and 19th of October.
- UseGalaxy.eu Upgraded to 19.09
Platforms that were referenced at least twice in the past month’s publications:
26 : CPT 16 : Huttenhower 9 : RepeatExplorer 5 : UseGalaxy.eu 4 : Workflow4Metabolomics 3 : deepTools 2 : AmrPlusPlus 2 : ARGs-OAP 2 : DDBJ 2 : Pasteur 2 : PhenoMeNal 2 : RiboGalaxy 2 : UseGalaxy.org.au
Note: This is the first time, since the beginning of time, that the Huttenhower server was not the most referenced server in the previous month.
Doc, Hub, and Training Updates

By [Simon Bray](https://training.galaxyproject.org/training-material/hall-of-fame#simonbray). In this tutorial, you will perform docking of ligands into the N-terminus of Hsp90 (heat shock protein 90). The tools used for docking are based on the open-source software [AutoDock Vina](http://vina.scripps.edu/) ([Trott and Olson 2009](https://galaxyproject.github.io/training-material/topics/computational-chemistry/tutorials/cheminformatics/tutorial.html#Trott2009)).

By [Graham Etherington](https://training.galaxyproject.org/training-material/hall-of-fame#ethering) and [Nicola Soranzo](https://training.galaxyproject.org/training-material/hall-of-fame#nsoranzo). Learn to use scater ([McCarthy et al. 2017](https://training.galaxyproject.org/training-material/topics/transcriptomics/tutorials/scrna-scater-qc/tutorial.html#10.1093/bioinformatics/btw777)) to visualise scRNA-seq data, filter out low-quality cells, confirm that filtering has worked, and then look at confounding factors such as batch effect to see if the data is biased to any technical artifacts.

By [Simon Gladman](https://training.galaxyproject.org/training-material/hall-of-fame#slugger70), [Helena Rasche](https://training.galaxyproject.org/training-material/hall-of-fame#erasche). This tutorial outlines the process to get your data out of Galaxy and to delete it from Galaxy afterwards.
Who’s Hiring
The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back! Have a Galaxy-related opening? Send it to outreach @ galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Full-Time, Rixensart, Belgium.
Develop core bioinformatics pipelines using a variety of environments (incl. Linux, Galaxy, DNA-Nexus) for base-calling, demultiplexing, data quality control, sequence alignment, variant calling, quantifying gene expression…

The Plant Computational Genomics lab in the Department of Ecology and Evolutionary Biology at the University of Connecticut seeks motivated MS and PhD students to join the lab in the Summer/Fall 2020.

AbSci, Vancouver, Washington, United States

Interested in an exciting new Galaxy-focused opportunity with core team members? Contact recruiting@galactic-core.com today.

The The European Galaxy Team has open positions in Freiburg, Germany.

Releases

Tripal Galaxy is a Drupal module is designed to support integration of the Tripal online database construction toolkit with Galaxy. It uses the blend4php library.
Release 1.1 features bug fixes, simplified results pages, and the results list now honors the Galaxy workflow star icons that indicates when intermediate files should be hidden. It’s cleaner and more simple.

A new version of the Galaxy Helm chart is available. The Galaxy Helm chart focuses on production-ready Galaxy deployments for Kubernetes that are easy to deploy and manage. Give it a try and help us make it better.

Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. These releases included numerous fixes and enhancements. See GitHub for details.
Other News
The GenomeSpace project ends on November 15, 2019 due to expiration of its NHGRI funding and we will be shutting down the GenomeSpace servers on that date. Galaxy was a GenomeSpace-enabled tool from the beginning of GenomeSpace and worked closely to maintain that connection over the years.


