September 2020 Galactic News
BCC2020 & GCC2021, events, platforms, blog posts, tutorials, pubs, releases, obs ...
On this page
In the September 2020 issue
- BCC2020 Wrap up and GCC2021 is on the way
- More event news
- Galaxy platform news
- Blog posts (lots of them)
- Training material and doc updates
- Publications (a boatload of them)
- Who’s hiring
- New releases
- Other news too
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
BCC2020 is Done; GCC2021 is Coming
The 2020 Bioinformatics Community Conference (BCC2020) brought together over 800 people for the second joint Bioinformatics Open Source Conference (BOSC) and Galaxy Community Conference (GCC). There are several write ups of the event:
- Birthdays and Online Conferences go Viral. ISMB2020 & BCC2020 in the time of COVID-19, a GIGABlog post by Chris Armit.
- Getting introduced to Bioinformatics and Open Science through BCC 2020, by Gigi Kenneth
- Lessons learnt from organizing a virtual conference (BCC2020), by BCC2020 Organizers

- 804 people registered for one or more events
- 592 for the three day meeting
- 443 for training
- 393 for CollaborationFest
- Training
- 60 training sessions
- covering 36 topics (18 with a Galaxy emphasis)
- adding up to 150 hours of live instruction
- Presentations
- 3 live keynotes (and then 3 recorded keynotes)
- 70+ talks, presented in both hemispheres
- 80+ posters and demos
- 20 Birds-of-a-Feather & 2 Socials

Videos of most presentations and training are now available on the conference web site. Training materials for many tutorials are also linked to.
Slides & Posters?
If you presented a talk or poster/demo then please upload your slides and PDFs to the F1000Research Galaxy Gateway. Once they are there, we will link to them from the conference site. (You can also send links/PDFs to outreach@galaxyproject.org.)

The 2021 Galaxy Community Conference (GCC2021) will be held in Ghent, Belgium, 5-12 July. We are hoping to be back in person next year. GCC2021 will end shortly before the The Ghent Festivities, 10 days of varied and free offer (music) performances, (street) theatre, exhibitions, animation for children, fairs, parades and much more.
Watch this space for details, and we hope to see you there.
More Event News
Despite COVID-19, there is still a lot going on. Some of it is virtual, but live events are starting to happen again, especially in Europe. We have updated our list of events to reflect what we know. Some highlights:

The next two roundtables are September 3 (this Thursday) and September 17. This week’s topics are interactive PR commands by Alex Mahmood, and workflow updates - format, tooling, and GUI by John Chilton.

Five workshops this fall, all in Jouy-en-Josas and in French, and starting September 10. Oui.

There are two upcoming Galaxy Australia workshops offered by QCIF on Galaxy (Intro and RNA-Seq Analysis). The first one is 8 September.

There are 15 events in the next 4 months
- 5 are online
- The remaining 10 are in France (6), Australia (2), and the United States (2)
And material from some recent events is now available:
- End user open source bioinformatics tools, by Peter van Heusden at Bioinformatics Hub of Kenya
- MPDS and Open Source Tools for Computer Aided Drug Discovery, by Lijo John, S Nagamani, Himakshi Sarma at DDH2020 Training Program
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. Here’s the recent platform news we know about:

PiRATE, a Pipeline to Retrieve and Annotate Transposable Elements, is a Galaxy VM for detecting, classifying and annotating TEs of non-model organisms. It has a tutorial and user support. PiRATE is supported by IFREMER.
The UseGalaxy.fr server is a general purpose omics analysis server hosted by the French Institute of Bioinformatics (IFB). In addition, this server also hosts the specialized Workflow4Metabolomics and ProteoRE servers.
User support is provided through a discourse-based forum.
A comprehensive set of data preprocessing, machine learning, deep learning and visualisation tools, consolidated workflows for end-to-end machine learning analysis and training materials to showcase the usage of these tools. This workbench includes scikit-learn, Keras (a deep learning library based on TensorFlow) and various other tools to transform, learn and predict and plot your data. The server is supported by the Goecks Lab at OHSU and the Freiburg Galaxy Project.
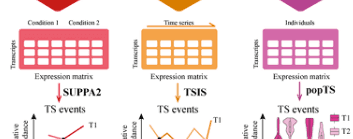
deepTS is a Galaxy platform for identifying, visualizing and analyzing Transcriptional Switch (TS) events from pairwise, temporal and population transcriptome data. A tutorial and user support are provided. deepTS is available as both a server and a Docker container. It is supported by Northwest A&F University.

BioCompute Objects and BioCompute Object Galaxy server supports the IEEE 2791-2020 standard for bioinformatics pipelines/workflows in the High-Throughput Sequencing (HTS) space. This effort is supported by the FDA and the HIVE Lab at George Washington University.

- Lots of tool updates on UseGalaxy.eu and UseGalaxy.org.au
- Galaxy Europe [is joining the NFDI-DataPLANT consortiu]m](https://galaxyproject.eu/posts/2020/08/22/DataPLANT/)
Platforms that were referenced/used at least twice in recent publications:
86 : Huttenhower 24 : UseGalaxy.eu 22 : RepeatExplorer 19 : ARGs-OAP 8 : UseGalaxy.org.au 7 : Galaxy-P 6 : Cistrome 5 : Pasteur 5 : Workflow4Metabolomics 4 : Globus Genomics 4 : Orione 4 : UseGalaxy.org 3 : ARGalaxy 3 : Mississippi 2 : APOSTL 2 : BioCiphers 2 : BioTeam 2 : DDBJ 2 : G-OnRamp 2 : GCAC 2 : Genomic Hyperbrowser 2 : GIO 2 : GVL-MEL 2 : ImmPort 2 : LAPPS Grid 2 : Osiris 2 : PhenoMeNal 2 : PreSTIGE 2 : RiboGalaxy 2 : SouthGreen 2 : SymD
Galactic Blog Activity
Here’s what people are saying right now:
By Nuwan Goonasekera, Alexandru Mahmoud, Mohamad Safadieh, Enis Afgan
A data browser, project-level isolation, and the latest release of Galaxy.

By StreetScience
Studying the RNA modifications of SARS-CoV-2 to improve understanding of SARS viruses.

By Anne Fouilloux
July has been a very busy month for Galaxy Climate ending with several sessions dedicated to Climate Science at BCC2020.

By Björn Grüning and Beatriz Serrano-Solano
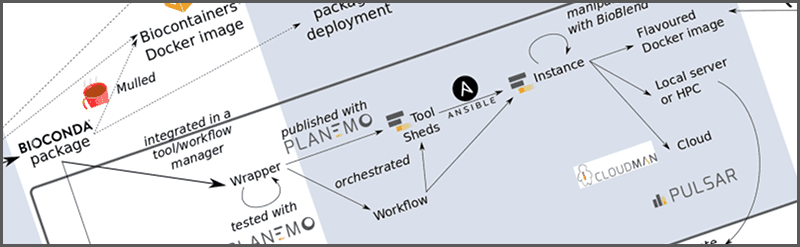
How you can get your software tool into a Galaxy server and thus, exposed to thousands of researchers. For this purpose, we will follow David Lopez’s steps to add the very generic UNIX diff tool to Galaxy.
By Peter van Heusden
The COVID-19 pandemic has forced a rapid - and unplanned - shift towards online events instead of the usual roster of face-to-face seminars and conferences.

By Dave Clements
Ten thousand publications have used, referenced, extended and implemented Galaxy in the past 15 years. Half of those pubs appeared in just the last 3 years. Learn about this and other trends in the Galaxy community, as revealed by what we are publishing.

By Delphine Lariviere, Sergei Kosakovsky Pond, Anton Nekrutenko + HyPhy and Galaxy Teams
An on-line, curated resource that serves two functions:
- Provide a coordinate conversion table between amino-acid coordinates in SARS-CoV-2 and a set of well-studied coronaviruses;
- Annotate amino acid residues of SARS-CoV-2 using available coronavirus literature.
Doc, Hub, and Training Updates
This tutorial was updated by Pavankumar Videm to use the most recent versions of tools. Learn how to use CLEAR-CLIP data in your analysis.

By Peter van Heusden, Simon Gladman, and Thoba Lose
A tutorial on variant calling in M. tuberculosis. This was first conceived at a workshop at NUBRI in Khartoum, Sudan, then presented at the African Society for Bioinformatics and Computational Biology meeting in Kumasi, Ghana in November 2019.
This tutorial was updated and expanded by Simon Bray. An introductory guide to using GROMACS (Abraham et al. 2015) in Galaxy to prepare and perform molecular dynamics on a small protein.

Sections on job configuration in Ansible were added to the Galaxy Installation with Ansible tutorial by Simon Gladman and to the Running Jobs on Remote Resources with Pulsar by Helena Rasche.

This tutorial saw a major revamp by Helena Rasche. Learn how to use Ansible for your deployment recipes.

The new GTN in Galaxy Webhook enables trainees to view training directly within Galaxy. This has been well received:
This is awesome! Magic @hexylena!
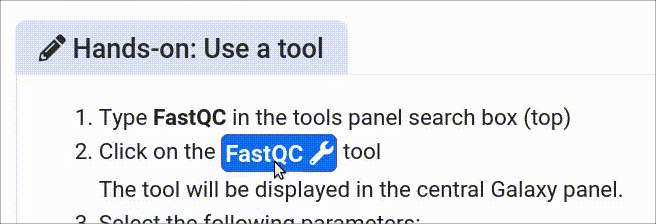
The GTN in Galaxy Webhook enables trainees to view training directly within Galaxy. As part of this, trainees can click on tools, and have those tools directly activated in Galaxy.

By Xavier Garnier, Anthony Bretaudeau, Anne Siegel, and Olivier Dameron
This new tutorial (and matching slides)uses data from the RNA-seq counts to genes tutorial in the AskOmics web application to do data integration and querying using the semantic web technologies

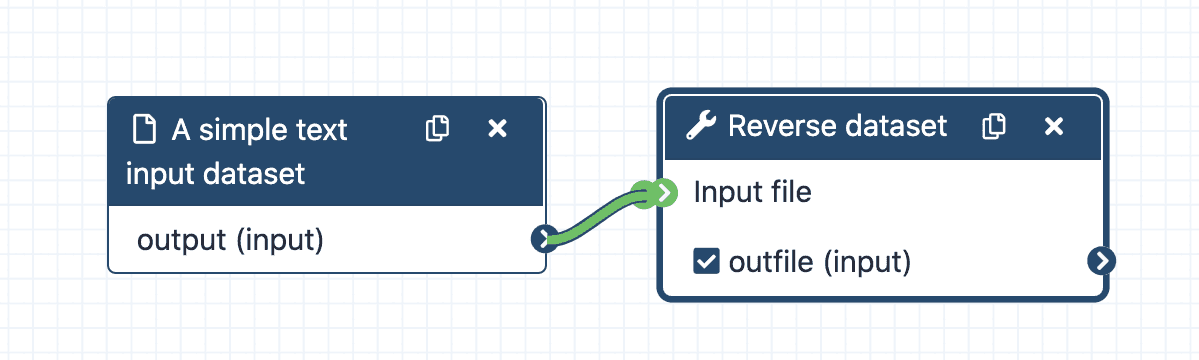
Learn how to use Galaxy’s Workflow Editor to construct different reusable workflows.
These slides were updated by by Nicola Soranzo to reflect the 14.0 release of BioBlend. BioBlend is a Python library for interacting with Galaxy programmatically.

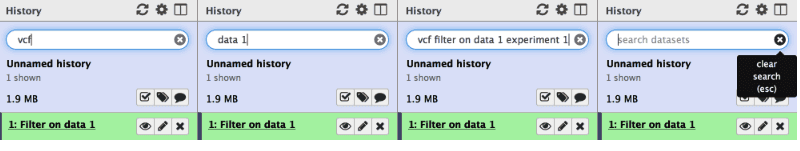
This tutorial got a major update from Helena Rasche. Feeling overwhelmed by all the work you have done in Galaxy? Don’t be. Use History search to find what you are looking for.
This slide deck and related resources got a thorough update from John Chilton. John also created several videos on this topic as well.
Ask and you shall receive. Thanks to work by Saskia Hiltemann and Helena Rasche there is now a GTN statistics page, listing all sorts of cool information about the state of the GTN library.

Publications
671 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in June, July and August. There were a boatload1 of Galactic and Stellar publications added, and a truckload1 of them are open access:
1 Who needs a Units of Measurement Ontology!
Suderman, K., Ide, N., Marc, V., Cochran, B., & Pustejovsky, J. (2020, June 30). ACL 2020 Workshop NLP-COVID.
Gu, Q., Kumar, A., Bray, S., Creason, A., Khanteymoori, A., Jalili, V., Grüning, B., & Goecks, J. (2020). BioRxiv, 2020.06.25.172445. doi: 10.1101/2020.06.25.172445
Rasche, H., & Hiltemann, S. (2020). GigaScience, 9(6). doi: 10.1093/gigascience/giaa065
Rasche, H., & Gruening, B. A. (2020). BioRxiv, 2020.08.23.263509. doi: 10.1101/2020.08.23.263509
Lopez-Delisle, L., Rabbani, L., Wolff, J., Bhardwaj, V., Backofen, R., Grüning, B., Ramírez, F., & Manke, T. (2020). Bioinformatics. doi: 10.1093/bioinformatics/btaa692
Tangaro, M. A., Donvito, G., Antonacci, M., Chiara, M., Mandreoli, P., Pesole, G., & Zambelli, F. (2020). GigaScience, 9(4). https://doi.org/10.1093/gigascience/giaa033
McGowan, T., Johnson, J. E., Kumar, P., Sajulga, R., Mehta, S., Jagtap, P. D., & Griffin, T. J. (2020). GigaScience, 9(4). https://doi.org/10.1093/gigascience/giaa025
Baker, D., Beek, M. van den, Blankenberg, D., Bouvier, D., Chilton, J., Coraor, N., Coppens, F., Eguinoa, I., Gladman, S., Grüning, B., Keener, N., Larivière, D., Lonie, A., Pond, S. K., Maier, W., Nekrutenko, A., Taylor, J., & Weaver, S. (2020). PLOS Pathogens, 16(8), e1008643. https://doi.org/10.1371/journal.ppat.1008643
Lac, M. du, Duigou, T., Herisson, J., Carbonell, P., Swainston, N., Zulkower, V., Shah, F., Faure, L., Mahdy, M., Soudier, P., & Faulon, J.-L. (2020). BioRxiv, 2020.06.14.145730. https://doi.org/10.1101/2020.06.14.145730
Sargent, L., Liu, Y., Leung, W., Mortimer, N. T., Lopatto, D., Goecks, J., & Elgin, S. C. R. (2020). PLOS Computational Biology, 16(6), e1007863. https://doi.org/10.1371/journal.pcbi.1007863
Jalili, V., Afgan, E., Gu, Q., Clements, D., Blankenberg, D., Goecks, J., Taylor, J., & Nekrutenko, A. (2020). Nucleic Acids Research, 48(W1), W395–W402. https://doi.org/10.1093/nar/gkaa434
Spoor, S., Wytko, C., Soto, B., Chen, M., Almsaeed, A., Condon, B., Herndon, N., Hough, H., Jung, S., Staton, M., Wegrzyn, J., Main, D., Feltus, F. A., & Ficklin, S. P. (2020). Database, 2020. https://doi.org/10.1093/database/baaa032
Sweeney, B. A., Tagmazian, A. A., Ribas, C. E., Finn, R. D., Bateman, A., & Petrov, A. I. (2020). Current Protocols in Bioinformatics, 71(1), e104. https://doi.org/10.1002/cpbi.104
Kancherla, J., Yang, Y., Chae, H., & Bravo, H. C. (2020). Bioinformatics. https://doi.org/10.1093/bioinformatics/btaa591
Wang, J., Alvin Chew, B. L., Lai, Y., Dong, H., Xu, L., Balamkundu, S., Cai, W. M., Cui, L., Liu, C. F., Fu, X.-Y., Lin, Z., Shi, P.-Y., Lu, T. K., Luo, D., Jaffrey, S. R., & Dedon, P. C. (2019). Nucleic Acids Research, 47(20), e130–e130. https://doi.org/10.1093/nar/gkz751
Silva, B. S. M. L., Heringer, P., Dias, G. B., Svartman, M., & Kuhn, G. C. S. (2019). PLOS ONE, 14(12), e0223466. https://doi.org/10.1371/journal.pone.0223466
McCann, J., Macas, J., Novák, P., Stuessy, T. F., Villaseñor, J. L., & Weiss-Schneeweiss, H. (2020). Frontiers in Plant Science, 11. https://doi.org/10.3389/fpls.2020.00362
Powell, G. L., Vannan, A., Bastle, R. M., Wilson, M. A., Dell’Orco, M., Perrone-Bizzozero, N. I., & Neisewander, J. L. (2020). Scientific Reports, 10(1), 11291. https://doi.org/10.1038/s41598-020-67966-8
Thursby, S.-J., Lobo, D. K., Pentieva, K., Zhang, S.-D., Irwin, R. E., & Walsh, C. P. (2020). GigaScience, 9(6). https://doi.org/10.1093/gigascience/giaa066
Publications are tagged with how they use, extend or reference Galaxy. This batch of pubs were tagged as:
448 : Methods 185 : UsePublic 78 : Workbench 77 : UseMain 42 : RefPublic 38 : Reproducibility 36 : UseLocal 28 : Tools 25 : IsGalaxy 12 : Education 9 : Shared 7 : Cloud 7 : Project 6 : Visualization 5 : Other 2 : HowTo 1 : Unknown 1 : UseCloud
Who’s Hiring
Quatre postes (CDI) de bioinformaticiens / biostatisticiens (H/F) à l’Institut Pasteur (Paris).

The Plant Computational Genomics laboratory at the University of Connecticut (Storrs, CT) has an opening for a Postdoctoral Scholar, developing mechanisms for cross-platform data/application sharing that builds on existing efforts with Galaxy, the Tripal platform, and cloud-based HPC.

The Schatz Lab at Johns Hopkins University is looking for
- self-driven individuals that can work independently to fill multiple software development positions on the Galaxy Project.
- ambitious individuals to fill a programmer analyst position working on the Galaxy and AnVIL projects.

The Barcelona Supercomputing Center - Centro Nacional de Supercomputación (BSC-CNS) is hiring.

VIB-UGent Center for Plant Systems Biology, Ghent, Belgium.
… We are building on the internationally used platform FAIRDOMhub for data management, and Galaxy (https://www.usegalaxy.be and https://usegalaxy.eu) for data analysis.

Black Canyon Consulting at NCBI, Bethesda, Maryland, United States.

Releases
Command-line utilities to help with managing users, data libraries and tools in a Galaxy instance, using the Galaxy API via the Bioblend library. This release features new commands (search toolsheds, delete users, uninstall tools), fixes & updates, and newly extended documentation.

BioBlend is a Python library for interacting with CloudMan and Galaxy‘s API. BioBlend makes it possible to script and automate the process of cloud infrastructure provisioning and scaling via CloudMan, and running of analyses via Galaxy.
See the release notes for what’s new in release 0.14.0.
Other news
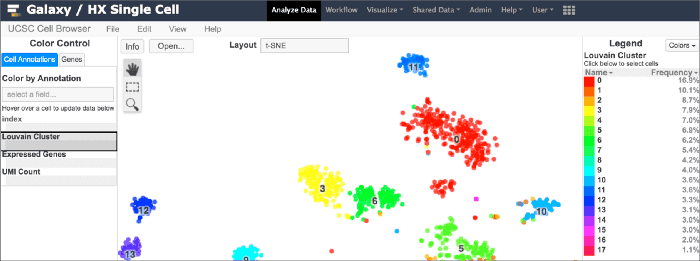
This write up by the Chan Zuckerberg Science Initiative includes a section on Irene Papatheodorou group’s work on the Human Cell Atlas.
Galaxy’s graphical workflow editor includes an automatic/default workflow layout algorithm that is 10 years old. A replacement based on elkjsis on the way, thanks to work by Helena Rasche.