Workflow Variables
On this page
Where are variables used?
Variables are used in the editor in the far right panel, under the section “Edit Step Actions”.
How to identify the input datasets variables?
The screenshot allows us to visualize that the Tophat2 tool is selected and configured to expect 2 .fastq files. In the far right panel, when we search for ‘Data input’, the input variables appear right in front of it between quotes. (eg. RNA-Seq FASTQ file, forward reads Data input ‘input1’ (fastqsanger)). Therefore the input variables for .fastq files are input1 and input2.
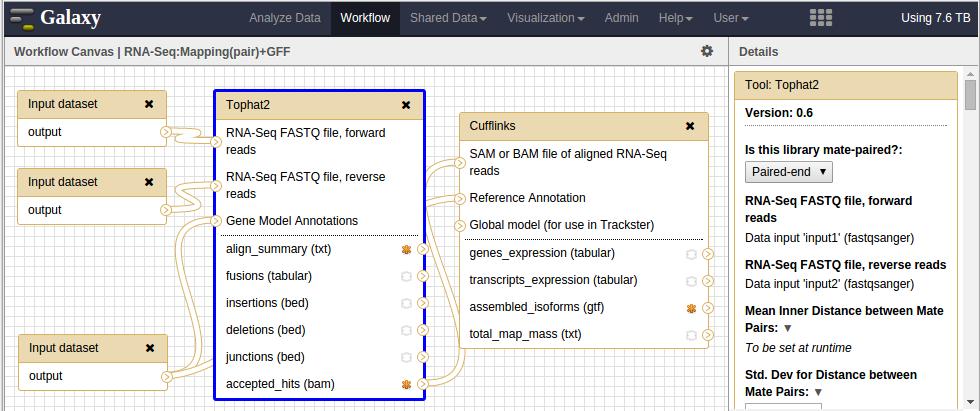 workflow editor showing the input variables names
workflow editor showing the input variables names
Usage
Variable syntax: #{input_name}
Full usage:
Proper syntax is: #{input_file_variable | option 1 | option n}
where
input_file_variable = is the name of an module input variable ("input_name" is supported)
| = the delimiter for added options. Optional if no options
options = basename, upper, lower
basename = keep all of the file name except the extension
(everything before the final ".")
upper = force the file name to upper case
lower = force the file name to lower case
Rename output based on specified content
When you run a tool, the result will be new boxes on the ‘History’ panel with names like ‘42 Tophat2 on data 27 and data 26: accepted_hits’. It can be hard to keep track if you’re running tens of samples using a workflow. You can rename the output based on the original input file name. In the section Edit Step Actions, you select ‘Rename Dataset’, choose the output and provide a new name. The new names can contain variables in order to tag the output. The screenshot shows the accepted_hits output being renamed to #{input_name}. For example, if your input filename is sample42.fastq the accepted_hits output will be named sample42.fastq.bam.
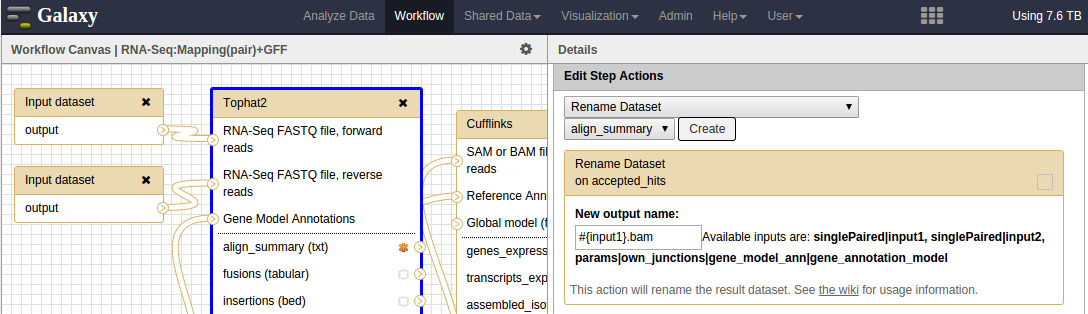 workflow editor showing the use of input variables in the renaming tool
workflow editor showing the use of input variables in the renaming tool
Source
https://github.com/galaxyproject/galaxy/blob/dev/lib/galaxy/jobs/actions/post.py
