March 2019 Galactic News
GalaxyAdmins, GCC2019 Childcare sponsors, Platforms, Pubs, Jobs, Blog!, Training, Tools, Releases and more!
On this page
The March 2019 Galactic News is here! This is a summary of what is going on in the Galaxy community.
- Event News
- eLife and F1000Research sponsor childcare at GCC2019
- The GalaxyAdmins meetup is back March 21!
- 184 new publications, great resources lead to great insight.
- Some most excellent Galaxy Platform News, including metagenomics, predictive model building, epigenome-wide association studies, immunogenomics, DNA methylation analysis, and more!
- An interview on The Galactic Blog with Nate Coraor about CVMFS & Galaxy.
- At least 15 Open positions in five countries on two continents.
- A burst of Training Updates.
- ToolShed contributions.
- Galaxy 19.01 released!
- And a bunch of other news too.
If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
GCC2019 Sponsors
We are very pleased to announce that eLife and F1000Research are first two sponsors for GCC2019. And they are both sponsoring childcare. Yes, there will be childcare at GCC2019!
eLife
eLife is a non-profit open access journal, publishing research from across the life and biomedical sciences. We also invest heavily in software development, new product design, collaboration and outreach. This includes the development of open-source tools, with extensible capabilities, that can be used, adopted and modified by any interested party to help move towards an ecosystem that serves science and scientists as efficiently and as cost-effectively as possible.
Have you got an idea that will bring cutting-edge technology to open research communication? We are here to help.
eLife’s Innovation Initiative was set up to identify and support innovative, small-to-medium-scale open-source projects aiming to improve the sharing and reuse of scientific discoveries. The primary outputs of the Initiative are open tools, technologies and processes aimed at improving the discovery, sharing, consumption and evaluation of important scientific research.
Through this Initiative, we’re always on the lookout for opportunities to engage with the best emerging talent and ideas at the interface of research and technology. You can find out more about some of these engagements on eLife Labs, or contact our Innovation Community Manager for more information (innovation@elifesciences.org).
The Galaxy Gateway on F1000Research
F1000Research is proud this year to be sponsoring childcare for GCC2019, which we feel is vital for inclusivity. F1000Research is an Open Access publishing platform for the life sciences that publishes a range of different article types, from traditional research articles, to method articles and software tools. The platform is particularly well suited to publishing software tool articles as our versioning system allows small updates at any time, we have proper syntax highlighting and we can support LaTeX and markdown submissions.
Our Galaxy Gateway on the platform brings together articles, posters and slides from Galaxy events. The Gateway allows authors to quickly share their work with the community, boosting visibility and giving researchers proper credit for their contribution.
We’re pleased to be running a call for papers for GCC2019 and would like to invite the Galaxy community to submit their articles to the Galaxy Gateway by 20th July 2019. These can be any type of article, such as a new Galaxy workflow or an update to an existing method or software tool. This is a great opportunity to shine a light on your research, improve its impact and be credited with an academic paper for your work. Please submit via the Galaxy gateway and if you have any questions at all do not hesitate to get in touch at publishers@f1000.com.
Your Organization/Vendor Here!
GCC2019 is looking for sponsors! If your organization wants to help put on the premier Galaxy event of the year, then please contact the organizers). And please encourage your vendors to consider sponsoring as well. Sponsors are potential partners for participants and their contributions make GCC affordable (and maybe even possible).
GalaxyAdmins
 ](/community/galaxy-admins/)
](/community/galaxy-admins/) 
After a two year hiatus, the online GalaxyAdmins meetups will return on March 21, with a community discussion, and a presentation from Pablo Moreno on Running Galaxy on Kubernetes, online at 15:00 GMT (see your local time). See the meetup page for links).
GalaxyAdmins is a discussion group for Galaxy community members who are responsible for Galaxy installations. Our online meetups are around an hour long and feature a presentation followed by an open discussion. It’s a great place to catch up on what your fellow admins are thinking about.
Upcoming events
These and other Galaxy related events are coming up:
Publications
184 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in February.
Highlighted Publications
Galactic and Stellar publications from February:
- Développement d’outils et de méthodes pour l’analyse de données épigénomiques, Jonathan Laperle, M. Sc thesis, Université de Sherbrooke, 2019
- GCAC: galaxy workflow system for predictive model building for virtual screening, Deepak R. Bharti, Anmol J. Hemrom and Andrew M. Lynn. BMC Bioinformatics 2019 19 (Suppl 13) :550 doi:10.1186/s12859-018-2492-8
- EWAS-Galaxy: a tools suite for population epigeneticsintegrated into Galaxy, Katarzyna Murat, Björn Grüning, Paulina Wiktoria Poterlowicz, Gillian Westgate, Desmond J Tobin, *Krzysztof Poterlowicz. bioRxiv 553784; doi:doi.org/10.1101/553784 ”
- Statistical Mitogenome Assembly with Repeats, Fahad Alqahtani and Ion I. Mandoiu. 2018 IEEE 8th International Conference on Computational Advances in Bio and Medical Sciences (ICCABS)
- SMAGEXP: a galaxy tool suite for transcriptomics data meta-analysis, Samuel Blanck and Guillemette Marot. GigaScience, Volume 8, Issue 2, 1 February 2019, giy167, doi:10.1093/gigascience/giy167
- Enrichment of phosphate solubilizing bacteria during late developmental stages of eggplant (Solanum melongena L.), Huixiu Li, Xiaoyan Ding, Chen Chen, Xiangnan Zheng, Hui Han, Chennan Li, Jingyang Gong, Ting Xu, Qing X Li, Guo-chun Ding, Ji Li. FEMS Microbiology Ecology, fiz023, doi:10.1093/femsec/fiz023
- Optimized Storing of Workflow Outputs through Mining Association Rules, Debasish Chakroborti, Manishankar Mondal, Banani Roy, Chanchal K. Roy, Kevin A. Schneider. 2018 IEEE International Conference on Big Data (Big Data), doi:10.1109/BigData.2018.8622351
- Biomolecular Reaction & Interaction Dynamics Global Environment (BRIDGE), Tharindu Senapathi, Simon Bray, Christopher B. Barnett, Björn Grüning, Kevin J Naidoo. Bioinformatics, btz107, doi:10.1093/bioinformatics/btz107
Five of the eight highlighted publications are open access.
Publication Topics
New Tag: Education
We added a new tag to the Galaxy Publication Library this month: Education. This tag highlights papers that focus on training, teaching, and/or education in general. Galaxy is well suited for teaching bioinformatics and this has been reflected in publications for several years. There are three new pubs this month about education. We also selectively back-curated several papers. Education is the first new tag added to the library in six years.
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 124 | +Methods | 50 | +UsePublic | 27 | +Workbench | 26 | +UseMain | |||
| 16 | +UseLocal | 12 | +RefPublic | 8 | +Tools | 7 | +IsGalaxy | |||
| 7 | +Reproducibility | 4 | +Cloud | 3 | +Education | 2 | +HowTo | |||
| 2 | +Shared | 1 | +Other |
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. A lot was added in February:
FreeBioInfo
FreeBioinfo is a public server for metagenomic analysis. The server supports a wide range of metagenomics and other tools. Ther is a Cookbook for the site (although it is in Chinese). FreeBioInfo is hosted at the China Agricultural University, in Beijing, China.
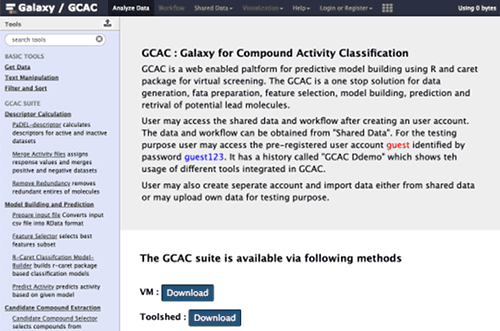
GCAC
GCAC : Galaxy for Compound Activity Classification is available as a public server, as a virtual machine, and as a tool suite in the Galaxy Toolshed. GCAC is a web enabled paltform for predictive model building using R and caret package for virtual screening. GCAC is a one stop solution for data generation, data preparation, feature selection, model building, prediction and retrieval of potential lead molecules. The GCAC Manual describes how to use the service. GCAC is supported by Jawaharlal Nehru University.
EWAS-Galaxy
EWAS-Galaxy is a Docker image for population epigenetics data analysis. Epigenome-wide association studies (EWAS) analyse genome-wide activity of epigenetic marks in cohorts of different individuals to find associations between epigenetic variation and phenotype. EWAS-Galaxy is supported by the University of Bradford.
Mandoiu Lab
The Mandoiu Lab Galaxy server is a customized installation of the Galaxy framework deploying bioinformatic tools and pipelines developed in the Mandoiu Lab at the University of Connecticut for high-throughput sequencing and immunogenomics data analysis. Source code and relevant publications for these and other tools developed in the lab are available on the Mandoiu lab software page.
DocMethyl
DocMethyl EpiMOLAS is Docker image for DNA methylation analysis in WGBS. This Galaxy docker container have tested with demo dataset under the Ubuntu 16.04 64 bit system equipped with four-core CPU, 32GB RAM, 400GB storage. The image is maintained by the Institute of Information Science, Academia Sinica, in Taipei, Taiwan.
Trinity

Trinity CTAT Galaxy is a public server and Docker container providing users with access to large memory resources required for de novo transcriptome assembly and downstream analysis. Funded by the National Cancer Institute, the Trinity Cancer Transcriptome Analysis Toolkit provides a suite of software for the analysis of cancer transcriptome data. The nature of cancer data requires different assumptions and approaches to analysis, above and beyond the normal challenges of dealing with RNA-seq. View more information on the tools available here. The resource has slides available for tasks such as moving data, and help can be reached at trinity@ncgas.org. There are no quotas, however, there is an aggressive purge policy for the disk usage. Files over 60 days old are removed by the system. An account is needed to access the resource and users are asked to fill out a registration form about your research (this helps us stay funded).
Galaxy Platforms in Publications
We tag papers that use, mention, implement or extend public Galaxy platforms (servers, services, clouds, containers…). Here are the counts for the past month’s publications:
New Galactic Blog Post
This month we have a Galactic Blog post where Nate Coraor was interviewed about what CVMFS actually is and what it does for Galaxy.
Who’s Hiring

The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back!
- Cloud Engineer V, Fred Hutchinson Cancer Research Center, Seattle, Washington, United States.
- ELIXIR Belgium has four Galaxy-related openings in Ghent:
- ELIXIR Open Science Community Manager, VIB-UGent Center for Plant Systems Biology
- ELIXIR Software developer data management tools, VIB-UGent Center for Plant Systems Biology
- ELIXIR Bioinformatics Trainer, VIB Bioinformatics Core
- ELIXIR Scientific Cloud Coordinator, VIB-UGent Center for Plant Systems Biology
- Software Developer, Harvard T.H. Chan School of Public Health, Boston, Massachusetts, United States. “Basic automated analysis workflows using Galaxy for 16S marker gene, metagenomic, and metatranscriptomic data leveraging existing software.”
- Cloud Computing Bioinformatics Programmer working with IRIDA, Simon Fraser University, Vancouver, Canada
- Senior IT DevOps Data Engineer and Senior IT Data Engineer (IT Bioinformatician), Adaptimmune, Abingdon, Oxfordshire, United Kingdom. Required: Experience with following bioinformatic pipeline tools: Galaxy…
- Specialist Solutions Architect — Biological Sciences, Amazon Web Services, United States
- Principal Technical Business Development Manager: AWS Research in Biomedical, Amazon Web Services, United States
- Bioinformatics Scientist, ResearchDx, Irvine, California, United States
- NGS Scientific Applications Specialist, Integrated DNA technologies, Iowa City, Iowa, United States
- The The European Galaxy Team has open positions, Freiburg, Germany
- Software engineer, system analysts/administrators, data analyst, and a comunnity and/or research manager
- The Blankenberg Lab in the Genomic Medicine Institute at the Cleveland Clinic Lerner Research Institute is hiring postdocs.
- Galaxy Project is hiring two software engineers at Johns Hopkins University, Baltimore, Maryland, United States.
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Doc, Hub, and Training Updates
Updates from the Galaxy Training Materials in February, and the Hub as well:
GTN Training Materials
- New Plates, Batches, and Barcodes slides by Mehmet Tekman.
- New Understanding Barcodes tutorial by Mehmet Tekman.
- New Mass spectrometry imaging: Examining the spatial distribution of analytes tutorial by Melanie Föll.
- New Pre-processing of Single-Cell RNA Data tutorial by Mehmet Tekman.
- Addition of data-library.yaml file for describing sample data of tutorials by Bert Droesbeke.
- Update to Visualization of RNA-Seq results with CummeRbund by Pavankumar Videm.
- Update to Making sense of a newly assembled genome by Ekaterina Polkh.
- Updates to Reference-based RNA-Seq data analysis by Andrea Bagnacani.
- Update to De Bruijn Graph Assembly, by Ekaterina Polkh.
- Update to CLIP-Seq data analysis from pre-processing to motif detection by Florian-H-Lab.
- Update to GO Enrichment Analysis by Daniel Sobral.
- Updates to Mapping and molecular identification of phenotype-causing mutations and Analysis of metagenomics data - The global picture by Anna Syme.
There was a tremendous amount of content added to the GTN while our fearless instructors of the 2019 Galaxy Admin Training faced below freezing temperatures in the unforgiving mountains of Pennsylvania. A warm thanks to Helena Rasche, John Chilton, Marten Cech, Nate Coraor, and Simon Gladman for adding to the growing Galaxy Server Administration section of the GTN. See these training materials below:
- New Ansible for installing Galaxy tutorial.
- New Galaxy Tool Management tutorial and slides.
- New Reference Data with CVMFS tutorial.
- New Upstream Authentication tutorial.
- New Empathy for admins slides.
- Update to Docker and Galaxy slides.
- Update to Connecting Galaxy to a compute cluster tutorial and slides.
- Update to Server Maintenance slides.
- Update to Galaxy Reports tutorial.
- Update to External Authentication slides.
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in February 2019 (there were a lot).
Releases
New additions to the Galaxy Ecosystem.
Galaxy Release 19.01
We are pleased to announce the release of Galaxy 19.01. This release includes the revised UI style, addition of colorful tags, extensive workflow enhancements, and enhanced support for Singularity.
Please see the release notes for additional information.
galaxy-lib 19.5.0
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
Other News
- From QCIF, both highlighting UseGalaxy.org.au
- We got an early look at the future of Galaxy management with Kubernates from Enis Afgan, stay tuned friends.