April 2019 Galactic News
GCC2019 registration and abstract submission, Platforms, Pubs, Jobs, Blog!, Training, Tools, Releases and more!
On this page
The April 2019 Galactic News is here! This is a summary of what is going on in the Galaxy community.
- Event News
- 120 new publications, great resources lead to great insight.
- Some most excellent Galaxy Platform News, including ways to investigate unmapped RNA-seq reads, language analysis, and RNA structure tools!
- A new entry to The Galactic Blog, about the upcomming GCC.
- At least 9 Open positions in three countries on two continents.
- Updates to training materials.
- ToolShed contributions.
- CloudBridge 2.0 released.
- Galaxy status page is live!
- And a bunch of other news too.
If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
GCC2019 Registration & Abstract Submission
We are pleased to announce that registration and abstract submission for the 2019 Galaxy Community Conference (GCC2019) are now open.
GCC2019 will be held 1-6 July in Freiburg, Germany. The tenth GCC will have many familiar features from earlier years, including accepted and lightning talks, posters and demos, birds-of-a-feather gatherings (BoFs), training, and a CollaborationFest. GCC2019 brings the most significant conference program update in several years:
- some training will be integrated with the main conference,
- three days of conference instead of two,
- several parallel sessions,
- and a strong emphasis on organizing content by domain.
GCC2019, like every GCC before it, will be built around community. Training topics are nominated, selected, and presented by the community. Presentations will feature the full spectrum of Galaxy applications, enhancements and deployments from the community as well.
If you are working in data intensive life science research then there will not be a better place to share your work, learn from others, and find new collaborators.
Please find additional information on GCC2019 here.
GCC2019 Sponsors
We are very pleased to welcome Limagrain as a Silver Sponsor for GCC2019.
Limagrain
As the fourth largest seed company worldwide, Limagrain’s mission is to move agriculture forward to meet global food related challenges. Limagrain is a cooperative group founded and managed by French farmers. Its parent company, Coopérative Limagrain, brings together nearly 2,000 farmers located in the center of France, in the Limagne Val d’Allier plain. The Group creates, produces and distributes field seeds, vegetable seeds and cereal products. Limagrain is present in 56 countries and has more than 10,000 employees. It makes nearly 2.5 billion Euros of sales with recognized brands on their markets: LG, Vilmorin, Hazera, Harris Moran, Jacquet, Brossard.
After participating for several years, Limagrain has the pleasure to help out with the Galaxy Community Conference organization for the first time ever. With over 200 Limagrain users worldwide and hundreds of tools dedicated to agronomic research, Galaxy has become central to Bioinformatics and Bioanalysis platforms of Limagrain research activities. By leveraging access to sequencing data of the plant species and by gathering tools and workflows tailored towards breeding, Galaxy is key to Limagrain competitiveness.
Your Organization/Vendor Here!
GCC2019 is looking for sponsors! If your organization wants to help put on the premier Galaxy event of the year, then please contact the organizers). And please encourage your vendors to consider sponsoring as well. Sponsors are potential partners for participants and their contributions make GCC affordable (and maybe even possible).
Upcoming events
These and other Galaxy related events are coming up:
Publications
120 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in March.
Highlighted Publications
Recently added Galactic and Stellar publications:
- GraphClust2: annotation and discovery of structured RNAs with scalable and accessible integrative clustering, Milad Miladi, Eteri Sokhoyan, Torsten Houwaart, Steffen Heyne, Fabrizio Costa, Björn Grüning, Rolf Backofen. bioRxiv 550335; doi: 10.1101/550335
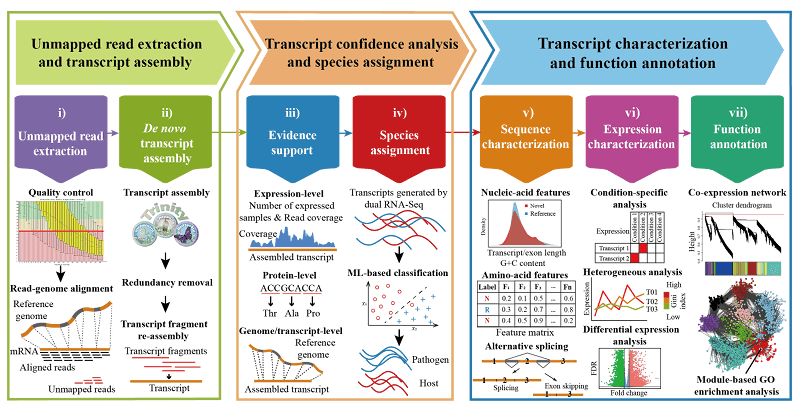
- CAFU: a Galaxy framework for exploring unmapped RNA-Seq data, Siyuan Chen, Chengzhi Ren, Jingjing Zhai, Jiantao Yu, Xuyang Zhao, Zelong Li, Ting Zhang, Wenlong Ma, Zhaoxue Han, Chuang Ma. Briefings in Bioinformatics, bbz018,doi:10.1093/bib/bbz018. 28 February 2019
- Visualization to Support Advanced Analysis of Genomic Data, Kumar Aman. Master Thesis, University of Oslo
- Designing an In Silico Strategy to Select Tissue-Leakage Biomarkers Using the Galaxy Framework, Nguyen L., Brun V., Combes F., Loux V., Vandenbrouck Y. (2019) In: Proteomics for Biomarker Discovery. Methods in Molecular Biology, Brun V., Couté Y. (eds), vol 1959.
Three of the four highlighted publications are open access.
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 83 | +Methods | 27 | +UsePublic | 16 | +UseMain | 14 | +Workbench | |||
| 11 | +UseLocal | 10 | +RefPublic | 6 | +Reproducibility | 5 | +IsGalaxy | |||
| 3 | +Tools | 1 | +Cloud | 1 | +Other | 1 | +Shared | |||
| 1 | +Visualization |
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. A lot was added in March:
GraphClust
The GraphClust2 server is a workflow for scalable clustering of RNAs based on sequence and secondary structures feature. GraphClust2 is implemented within the Galaxy framework and consists a set of integrated Galaxy tools and flavors of the linear-time clustering workflow.
Language Analysis Portal
The language analysis portal (LAP) provides an intuitive and easy-to-use web interface to a centralized repository of a wide range of language technology tools, all installed on a high-performance computing (HPC) cluster. Users are able to compose and run workflows using an in-browser graphical interface, with multiple tools and resources chained together in potentially complex pipelines. Although the project reaches out to a diverse set of user groups, it particularly aims to facilitate use of language analysis in the social sciences, humanities, and other fields without strong computational traditions.
CAFU
CAFU is a Galaxy-based bioinformatics framework for comprehensive assembly and functional annotation of unmapped RNA-seq data from single- and mixed-species samples which integrates plenty of existing NGS analytical tools and our developed programs, and features an easy-to-use interface to manage, manipulate and most importantly, explore large-scale unmapped reads.
Galaxy Platforms in Publications
We tag papers that use, mention, implement or extend public Galaxy platforms (servers, services, clouds, containers…). Here are the counts for the past month’s publications:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 11 | >Huttenhower | 5 | >Workflow4Metabolomics | 4 | >ARGs-OAP | 4 | >RepeatExplorer | |||
| 2 | >Langille | 2 | >UseGalaxy.eu | 1 | >AmrPlusPlus | 1 | >ARIES | |||
| 1 | >CAFU | 1 | >Cistrome | 1 | >CPT | 1 | >EGI | |||
| 1 | >Galaxy-P | 1 | [>Genomic Hyperbrowser](https://www.zotero.org/groups/1732893/galaxy/tags/>Genomic Hyperbrowser) | 1 | >GraphClust | 1 | >MPDSDM | |||
| 1 | >MPDSTB | 1 | >Pasteur | 1 | >PreSTIGE | 1 | >ProteoRE | |||
| 1 | [>RNA Workbench](https://www.zotero.org/groups/1732893/galaxy/tags/>RNA Workbench) | 1 | >SouthGreen |
New Galactic Blog Post
This month we have a Galactic Blog post where Björn Grüning gives a sneak peak at GCC2019.
Who’s Hiring

The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back!
- Scientist - Molecular R&D, MilliporeSigma, Rockville, Maryland, United States
- Biostatistician, Emory University, Atlanta, Georgia, United States
- Cloud Architect, Fred Hutchinson Cancer Research Center, Seattle, Washington, United States.
- ELIXIR Belgium has four Galaxy-related openings in Ghent:
- ELIXIR Open Science Community Manager, VIB-UGent Center for Plant Systems Biology
- ELIXIR Software developer data management tools, VIB-UGent Center for Plant Systems Biology
- ELIXIR Bioinformatics Trainer, VIB Bioinformatics Core
- ELIXIR Scientific Cloud Coordinator, VIB-UGent Center for Plant Systems Biology
- Software Developer, Harvard T.H. Chan School of Public Health, Boston, Massachusetts, United States. “Basic automated analysis workflows using Galaxy for 16S marker gene, metagenomic, and metatranscriptomic data leveraging existing software.”
- The The European Galaxy Team has open positions, Freiburg, Germany
- Software engineer, system analysts/administrators, data analyst, and a comnunity and/or research manager
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Doc, Hub, and Training Updates
Updates from the Galaxy Training Materials in March:
GTN Training Materials
- New Group tags for complex experimental designs mini-tutorial by Marius van den Beek.
- New Machine learning: classification and regression tutorial by Anup Kumar and Bérénice Batut.
- New Identification of somatic and germline variants from tumor and normal sample pairs tutorial by Wolfgang Maier.
- New Downstream single-cell RNA analysis with RaceID tutorial by Mehmet Tekman.
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in March 2019.
Releases
New additions to the Galaxy Ecosystem.
CloudBridge 2.0 release
CloudBridge is a Python library that offers a uniform interface to major Infrastructure-as-a-Service (IaaS) cloud providers. CloudBridge ensures operational consistency across the supported providers allowing the same code to run consistently across multiple cloud providers. With this release, CloudBridge supports Amazon Web Service (AWS), Microsoft Azure, OpenStack, and now Goodle cloud. This release is further characterized by improved code consistency.
Please see the release notes for additional information.
Galaxy Status
We are happy to announce that we now have a status page to monitor the status of our public useGalaxy.* servers and certain Galaxy services. This page tracks the operational status of usegalaxy.org, usegalaxy.eu, and usegalaxy.org.au servers, as well as Galaxy services including the Galaxy (Main) Tool Shed, Galaxy CloudLaunch, and Galaxy test services. Status.galaxyproject.org will serve as the central channel for announcing and tracking planned downtime and routine maintenance.
galaxy-lib 19.5.1
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
Other News
- Galaxy + Carperntries == Gallantries! A cross community training effort to combine the best of both worlds. See the announcement for additional details.
- Biohackathon2019 Call for Hacking projects is officially open. Apply by 7 April.