December 2019 Galactic News
Events, Pubs, Blogs, Platforms, Tutorials, Doc, Jobs, Releases, and Platypi too.
On this page
The December 2019 Galactic News is out:
- 12 Upcoming Events
- India, PAG, HTS, single cell/microbiome, Barcelona, and more
- 191 new publications
- Two new blog posts
- g:Profiler, GalaxyP
- Galaxy Platform News
- Systems biology, Laniakea, OpenRefine, …
- Training material and doc updates
- defining data types, citing GTN, Windows, account help, metatranscriptomics, Galaxy for everyone
- Openings
- with 5 organizations in Denmark, France, US, US, Germany
- Releases
- Galaxy 19.09 is out
- And other news too
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events

Galaxy (especially Galaxy Australia will be featured in a workshop and a plenary talk at the International Symposium on Bioinformatics 2019 (InSyB2019) being held December 21-22, 2019 at Hans Raj Mahila Maha Vidyalaya, Jalandhar, Punjab, India. Registration and abstract submission are still open.

Galaxy will be at Plant and Animal Genome XXVII (PAG 2020), in San Diego, California, United States, January 11-15. This includes a hands-on Galaxy Workshop (highlighting the new Excellence in Breeding platform) and many talks and posters featuring Galaxy.
And there will be a GMOD Codefest before PAG and an NCBI Codeathon after PAG.

This week-long Galaxy workshop in Freiburg will introduce participants to high-throughput data analysis with Galaxy. Space is limit, and applications are competitive. Apply by 19 December.

Galaxy will be at 2020 ABRF meeting, in Palm Springs, California, United States, February 29 through March 3. This includes a full day hands-on Galaxy Workshop about using Galaxy with your single cell and microbiome data. ABRF is the annual conference for technology-enabled multidisciplinary research.
Registration is now open, but space is limited.

Save these dates! The next Galaxy Admin Training will be offered 2-6 March at the Barcelona Supercomputing Center.
This week-long hands-on training will feature what you need to know to set up your own production quality Galaxy server. Registration will open later this month.

There are
- 12 upcoming events
- on 3 continents, plus online
- in India, US, Belgium, Germany, France, Spain, and the UK.
Publications
191 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month. There were seven Galactic and Stellar publications added, and all seven of them are open access:
Kumar, A., Grüning, B., & Backofen, R. (2019). BioRxiv, 838599. doi: 10.1101/838599
Wibberg, D., Batut, B., Belmann, P., Blom, J., Glöckner, F. O., Grüning, B., … Kohlbacher, O. (2019). F1000Research, 8, 1877. doi: 10.12688/f1000research.20244.1
Perez‐Riverol, Y., & Moreno, P. (2019). PROTEOMICS, 1900147. doi: 10.1002/pmic.201900147
Demko, M., Chrást, L., Dvořák, P., Damborský, J., & Šafránek, D. (2019). Microorganisms, 7(11), 553. doi:10.3390/microorganisms7110553
McGowan, T., Johnson, J. E., Kumar, P., Sajulga, R., Mehta, S., Jagtap, P. D., & Griffin, T. J. (2019). BioRxiv, 842856. doi: 10.1101/842856
Kumar, P., Johnson, J. E., Easterly, C., Mehta, S., Sajulga, R., Nunn, B., … Griffin, T. J. (2019). BioRxiv, 843078. doi: 10.1101/843078
Monnerie, S., Petera, M., Lyan, B., Gaudreau, P., Comte, B., & Pujos-Guillot, E. (2019). Metabolites, 9(11), 250. doi: 10.3390/metabo9110250
Publications are tagged with how they use, extend or reference Galaxy. The past month’s pubs were tagged as:
136 : +Methods 64 : +UsePublic 24 : +Workbench 19 : +UseLocal 19 : +UseMain 14 : +RefPublic 11 : +Tools 4 : +IsGalaxy 3 : +Reproducibility 2 : +Education 2 : +Unknown 2 : +Visualization 1 : +Shared
Galactic Blog Activity
Two new entries in the past month, both related to UseGalaxy.eu..
By Ivan Kuzmin. gProfiler is a toolset for functional enrichment analysis and conversions of gene lists.

By Ben Orsburn. The arguments are building up for why you need this.

Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. There are many new platforms this month:

The BioDivine server hosts tools for systems biology, developed by the Systems Biology Research Centre (Sybila) in the Faculty of Informatics of Masaryk University in Brno, Czechia.

Version 2 of Laniakea announced at the November ELIXIR Innovation and SME Forum. Laniakea provides automatic deployment of virtual Galaxy environments. See the video for details.
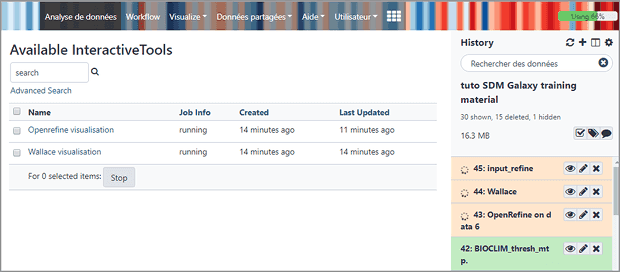
OpenRefine, a tool for cleaning data, is now available as an Interactive tool on Ecology Galaxy.

- UseGalaxy.eu: 10,000 users, 6,370,000 jobs and 11,900,000 datasets
- UseGalaxy.eu Tool Updates for 10th, 15th, 23rd, and 30th of November.
- Also see recent blog posts
Platforms that were referenced at least twice in the past month’s publications:
25 : Huttenhower 9 : UseGalaxy.eu 8 : CPT 8 : RepeatExplorer 5 : Workflow4Metabolomics 4 : Cistrome 4 : Galaxy-P 3 : UseGalaxy.org.au 2 : ARGs-OAP 2 : Orione
And the ProteoRE group is hiring too. See below.
Doc, Hub, and Training Updates
New comprehensive documentation by @selten on how to define data types for Galaxy.

Thanks to Helena Rasche, every GTN Tutorial now includes a Citing this Tutorial section, including a BibTex citation for the tutorial.

The documentation on how to run Galaxy on Windows has been completely revamped by Tomas Klingström to use the Windows Subsystem for Linux.

The account creation documentation was expanded by Jennifer Hillman-Jackson to cover additional situations and help resources.
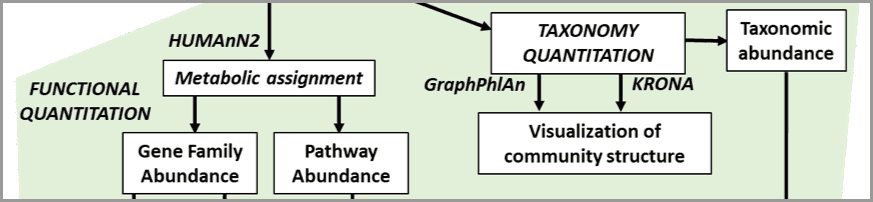
By Pratik Jagtap, Subina Mehta, Ray Sajulga, Bérénice Batut, Emma Leith, Praveen Kumar
Metatranscriptomics analysis examines how the microbiome responds to the environment by studying the functional analysis of genes expressed by the microbiome. This tutorial uses the ASaiM workflow (Batut et al. 2018) to study a time-series analysis of a microbial community from (Kunath et al. 2018).
By Anne Fouilloux, Nadia Goué, Christopher Barnett, Michele Maroni, Olha Nahorna, Dave Clements, Saskia Hiltemann. Familiarize yourself with Galaxy, even if you aren’t working in life sciences.

Who’s Hiring
Hey Good People! It’s hard to find good people, such as yourself, but surely you know good people who could help the good people below fill their open positions. (And if you have a Galaxy-related opening then please send it to outreach @ galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.)
Biotech Research & Innovation Centre (BRIC) Bioinformatics Core Facility, University of Copenhagen.
Maintain, update, upgrade and provide user support for a local version of a Galaxy server. (And other stuff too.) Applications close 15 December.

CEA/INSERM/UGA, Grenoble, France. Conception d’un workflow protéogénomiques: identifiez, optimisez, et le cas échéant, développez les outils necessaires dans Galaxy; ils seront déployés sur le serveur ProteoRE. Idéalement, le workflow sera documenté sous la forme de leçons intégrées dans le “Galaxy Training Network”.

The Plant Computational Genomics lab in the Department of Ecology and Evolutionary Biology at the University of Connecticut seeks motivated MS and PhD students to join the lab in the Summer/Fall 2020.

Take the Galaxy deployments to the next level with the core team members at Galactic Core. Email recruiting@galactic-core.com.

The The European Galaxy Team has open positions in Freiburg, Germany.

Releases

The 19.09 release of Galaxy is out and full of new features:
- Embedded reports in workflows
- New visualizations
- free style image annotation, allowing you to make notes on images in Galaxy and save these annotations (from Anup Kumar)
- OpenLayers based map visualization, allowing you to visualize your GIS data in Galaxy (also from Anup Kumar)
- And HyPhy-Vision (from iGEM/UCSD), and iCn3D
- Lots of new datatypes.
- A new type of interactive Galaxy tool has been implemented
- A New Toolshed Client Interface
Want the details? See the developer and admin release announcement and the user release announcement.
Other News
A US$3.5M NSF grant has been awarded to Penn State researchers to enable Galaxy-based tools to use GPUs. Speeding up science, by 10-100x.

The Galaxy Help online forum turned one year old November 19. This Discourse based site now has over 3500 posts and 1000 users. Give it a look.

And that is quite possibly the coolest end of year news item we have ever had.
Thanks for an excellent year, and looking forward to 2020.


