January 2020 Galactic News
Events, Pubs, Blogs, Platforms, Tutorials, Doc, and Happy New Year too
On this page
The January 2020 Galactic News is out:
- 13 Upcoming Events
- BCC2020 training topic nominations, Register for Galaxy Admin Training, Galaxy @ PAG & ABRF, and training @ Earlham and Rennes.
- 172 new publications
- Six highlighted pubs from RT-qPCR to functional genomics
- Two new blog posts
- PGP-UK and Crowdsourcing Science
- Galaxy Platform News
- Three new platforms, UseGalaxy.* news, and platforms in pubs
- Training material and doc updates
- R, RStudio, ATAC-Seq, Scanpy, and Single Cell RNA-Seq
- And other news too
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
Events
There is an unusually rich (and aromatic!) mix of upcoming Galaxy related events:

The Bioinformatics Community Conference (BCC2020) needs
- Nominate Training Topics by January 17
BCC is GCC + BOSC, and will be held in July in Toronto, right after ISMB 2020 in Montréal.

Registration is now open for the 2020 Galaxy Admin Training being offered 2-6 March at the Barcelona Supercomputing Center. This week-long hands-on training will feature what you need to know to set up your own production quality Galaxy server. Space is limited.

Galaxy will be at Plant and Animal Genome XXVII (PAG 2020), in San Diego, California, United States, January 11-15. This includes a hands-on Galaxy Workshop (highlighting the new Excellence in Breeding platform) and many talks and posters featuring Galaxy.
And there will be a GMOD Codefest before PAG and an NCBI Codeathon after PAG.

Galaxy will be at 2020 ABRF meeting, in Palm Springs, California, United States, February 29 through March 3. This includes a full day hands-on Galaxy Workshop about using Galaxy with your single cell and microbiome data. ABRF is the annual conference for technology-enabled multidisciplinary research.
Early registration ends January 15. Space is limited.

An introduction to Single-Cell Genomics featuring Galaxy. This course is for bench-based researchers planning a single-cell project. Registration deadline is 2 March.

Introduction à la manipulation de données de régions génomiques et à l’analyse de jeux de données de type NGS via Galaxy.

There are
- 13 upcoming events
- on 3 continents, plus online
- in US, Japan, Belgium, Germany, Spain, France, UK, and Canada.
Publications
172 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month. There were 9 Galactic and Stellar publications added, and 7 of them are open access. Some highlights:
Zanardi, N., Morini, M., Tangaro, M. A., Zambelli, F., Bosco, M. C., Varesio, L., … Cangelosi, D. (2019). Scientific Reports, 9(1), 1–12. doi: 10.1038/s41598-019-53155-9
Föll, M. C., Moritz, L., Wollmann, T., Stillger, M. N., Vockert, N., Werner, M., … Schilling, O. (2019). GigaScience, 8(12). doi: 10.1093/gigascience/giz143
Emami Khoonsari, P., Moreno, P., Bergmann, S., Burman, J., Capuccini, M., Carone, M., … Spjuth, O. (2019). Bioinformatics, 35(19), 3752–3760. doi: 10.1093/bioinformatics/btz160
Payá-Milans, M., Poza-Viejo, L., Martín-Uriz, P. S., Lara-Astiaso, D., Wilkinson, M. D., & Crevillén, P. (2019). GigaScience, 8(12). doi: 10.1093/gigascience/giz147
Etherington, G. J., Soranzo, N., Mohammed, S., Haerty, W., Davey, R. P., & Palma, F. D. (2019). A GigaScience, 8(12). doi: 10.1093/gigascience/giz144
Blum, M., Cholley, P.-E., Malysheva, V., Nicaise, S., Moehlin, J., Gronemeyer, H., & Mendoza-Parra, M. A. (2020). Life Science Alliance, 3(1). doi: 10.26508/lsa.201900546
Publication Topics
Publications are tagged with how they use, extend or reference Galaxy. The past month’s pubs were tagged as:
113 : +Methods 48 : +UsePublic 23 : +RefPublic 19 : +Workbench 17 : +UseMain 15 : +UseLocal 9 : +IsGalaxy 7 : +Reproducibility 6 : +Tools 3 : +Education 2 : +Cloud 2 : +Shared 1 : +Other 1 : +Visualization
Galactic Blog Activity
By Yvan Le Bras, Simon Bénateau.
Galaxy for Ecology, mixing Ecology research, Citizen Science and Massively Multi Online Science.

Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. There are many new platforms this month:
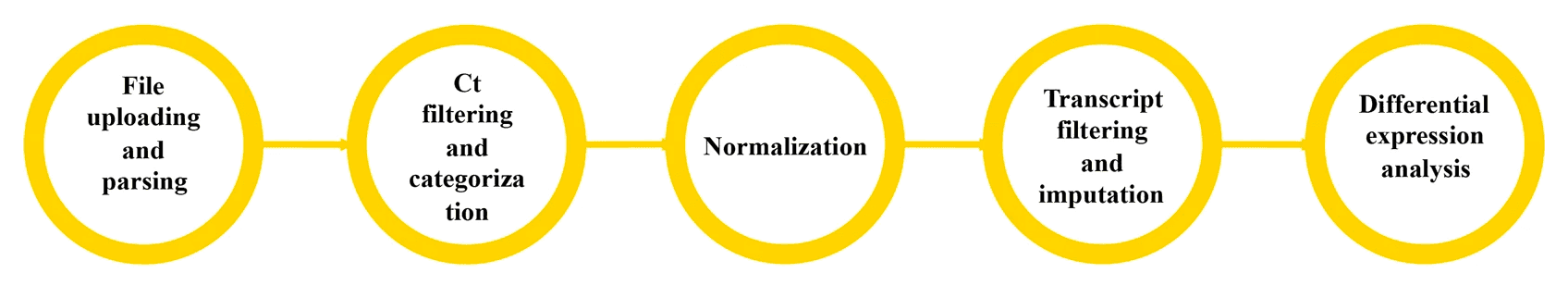

The GASLINI server supports PIPE-T, a tool for reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR) analysis. Requires an account, but anyone can create an account.

The Shapiro Lab at NCI provides RNA structure and design tools, including RNA2D3D and StructureLab, on a Galaxy virtual machine image.
This Galaxy docker instance contains tools and workflows aimed at the analysis of epigenomics data, both ChIP-Seq and RNA-Seq.
UseGalaxy.* News

- UseGalaxy.org.Au is expanding its range of tools to better support different communities including 19 metabolomics analysis tools & 2 workflows and 20 new tools for trimming, analysing, QC and assembling Oxford Nanopore data
- Galaxy Australia upgraded to Galaxy version 19.09
Galaxy Platforms in Publications
Platforms that were referenced at least twice in the past month’s publications:
25 : Huttenhower 8 : Workflow4Metabolomics 5 : UseGalaxy.eu 4 : CPT 4 : Galaxy-P 4 : RepeatExplorer 3 : ARGs-OAP 2 : Genomic Hyperbrowser 2 : GVL-Unspecified 2 : HiCExplorer 2 : Pasteur 2 : PhenoMeNal 2 : UseGalaxy.org.au
Doc, Hub, and Training Updates
Then the Galaxy Training Network wants your feedback. Please take a few minutes to let us know about your training experiences.

By Bérénice Batut, Fotis E. Psomopoulos, Toby Hodges.
We believe that every learner can achieve competency with R. Give it a try.

By Bérénice Batut, Fotis E. Psomopoulos, Toby Hodges.
This tutorial adapt the Carpentries “Intro to R and RStudio for Genomics” lesson to the Galaxy platform.
Lucille Delisle has updated the ATAC-Seq data analysis tutorial to add macs2 callpeak as an alternative to Genrich and addressed issues with bowtie indexes.
By Bérénice Batut.
Clustering single-cell data from 10x Genomics, including preprocessing, clustering and the identification of cell types via known marker genes, using Scanpy (Wolf et al. 2018).
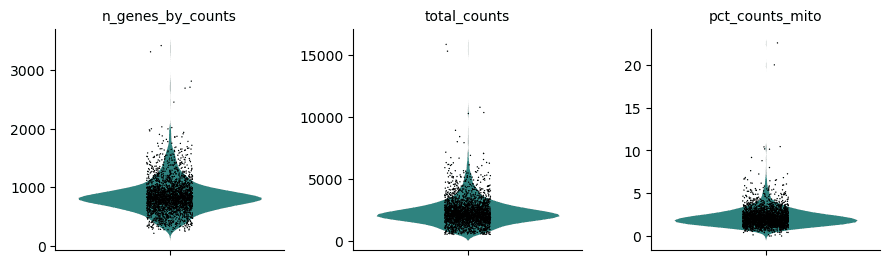
By Wendi Bacon, EMBL-EBI.
This webinar video highlights Galaxy interface for single cell analysis. Specifically it highlights Scanpy (which would otherwise require Python programming skills) to analyse a Drop-seq dataset located in EMBL-EBI’s Single Cell Expression Atlas.
Other News
Including a report about usegalaxy.eu and this year’s community conference in Freiburg.

Galaxy is a prominent part of the NCI ITCR Program that funds tools that support the analysis of –omics, imaging, and clinical data, as well as network biology and data standards.

Dr Chris Barnett has invested in the research cloud-computing platform ilifu to create a central resource where a like-minded research community can come together and share these tools and expertise, using the opensource platform Galaxy.



