July 2014 Galaxy Update
On this page
Welcome to the July 2014 Galaxy Update, a monthly summary of what is going on in the Galaxy community. Galaxy Updates complement the Galaxy Development News Briefs which accompany new Galaxy releases and focus on Galaxy code updates.
The Galaxy Update is going out a few days early this month because the usual release date is during GCC2014.
Events
GCC2014: June 30 - July 2, Baltimore
The 2014 Galaxy Community Conference (GCC2014) starts on Monday, June 30, and runs through July 2, at the Homewood Campus of Johns Hopkins University, in Baltimore, Maryland, United States. The program is online and all titles and abstracts for accepted talks and posters are now online.
Fifteen Training sessions on 12 topics, several Birds of a Feather, two lightning talk sessions, and the very first Galaxy Project Hackathon are also happening.
Galaxy @ ISBMB and BOSC 2014
There will be at least six talks and five posters related to Galaxy at ISMB and BOSC 2014 this year. Talks include
- Galaxy as an Extensible Job Execution Platform, John Chilton
- Enhancing the Galaxy Experience through Community Involvement, Daniel Blankenberg
- TT03: Interactive Visual Analysis with Galaxy Charts, Sam Guerler
- TT24: From the Ground to the Cloud in 25 minutes: Building a Customized Galaxy Analysis Server Using Only a Web Browser, Daniel Blankenberg
- TT27: Bioinformatics and Computer Biology Systems design applied to Medical Molecular Nanobiotechnology, Allan Orozco
- TT29: Scaling Galaxy: Preparing for Those Next Few Orders of Magnitude, John Chilton
Other Events
Over the rest of the summer there are other Galaxy related events in Leiden, Sydney, Brisbane, São Paulo, and Rio de Janeiro. Also see the Galaxy Events Google Calendar for details on other events of interest to the community.
New Papers
48 papers were added to the Galaxy CiteULike Group in June. Some papers that may be particularly interesting to the Galaxy community:
-
“BioBlend.objects: metacomputing with Galaxy”, by S. Leo, L. Pireddu, G. Cuccuru, et al. Bioinformatics (12 June 2014), doi:10.1093/bioinformatics/btu386
-
“RPPApipe: A pipeline for the analysis of reverse-phase protein array data” by Johannes Eichner, Yvonne Heubach, Manuel Ruff, et al. Biosystems (June 2014), doi:10.1016/j.biosystems.2014.06.009
-
“Using Bioinformatics Tools to Study the Role of microRNA in Cancer”, by Fabio Passetti, Natasha Andressa Nogueira Jorge, Alan Durham; In Clinical Bioinformatics, Vol. 1168 (2014), pp. 99-116, doi:10.1007/978-1-4939-0847-9_7
-
“Ocular and Extraocular Expression of Opsins in the Rhopalium of Tripedalia cystophora (Cnidaria: Cubozoa)” by Jan Bielecki, Alexander K. Zaharoff, Nicole Y. Leung, Anders Garm, Todd H. Oakley, PLoS ONE, Vol. 9, No. 6. (5 June 2014), e98870, doi:10.1371/journal.pone.0098870
The new papers were tagged in many different areas:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 2 | Cloud | 1 | Project | 3 | Tools | 4 | UsePublic | |||
| 2 | HowTo | 3 | RefPublic | - | UseCloud | - | Visualization | |||
| 1 | IsGalaxy | 1 | Reproducibility | 2 | UseLocal | 17 | Workbench | |||
| 16 | Methods | 1 | Shared | 7 | UseMain |
Who’s Hiring
The Galaxy is expanding! Please help it grow.
- Experimental Officer in Bioinformatics, NERC Metabolomics Facility, University of Birmingham, UK
- Two postdoc positions in integrative genomics available in Oslo, Norway
- Statistical Genomics Postdoc opening in the Makova lab at Penn State
- The Galaxy Project is hiring software engineers and post-docs
Got a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
New Public Servers
One new public Galaxy server was added to the published list in June:
GVL QLD
- Link: Genomics Virtual Lab GVL-QLD
- Domain/Purpose: General purpose Galaxy based on the Genomics Virtual Lab platform.
- Comments: Has 16 virtual CPUs.
- User Support:
- GVL Help
- Follow tutorials at GVL Learn and use Galaxy Tut
- Quotas:
- University of Queensland and collaborators: 2TB
- Other Australian Researchers: 1TB (make sure you register with your Institute email address)
- Other registered users: 200GB
- Unregistered users: 5GB
- Sponsor(s): Genomics Virtual Lab and the University of Queensland Research Computing Centre
Galaxy Distributions
June 2, 2014 Galaxy Distribution
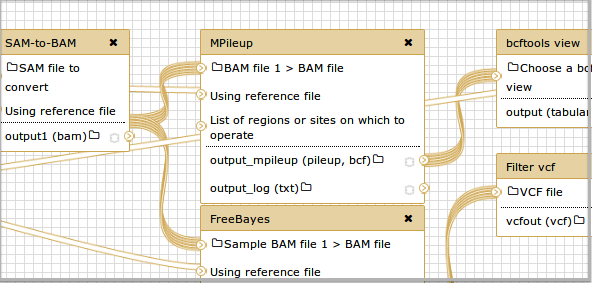
*example dataset collection workflow ([credits](/archive/dev-news-briefs/2014-06-02/#dataset-collections))*
**[News Brief](/archive/dev-news-briefs/2014-06-02/)** **Highlights:**
- Dataset Collections introduced
- Changes to database build (dbkey) organization
- Enhancements to Tool configuration and Workflow options
- Trackster, User Interface, and Admin panel upgrades
- Significant updates to Admin and Job functionality
- Tool Shed repository and API additions
- Data updates plus new Security features and Bug fixes
 |
getgalaxy.org | |
| galaxy-dist.readthedocs.org | ||
| bitbucket.org/galaxy/galaxy-dist | ||
| new: | $ hg clone https://bitbucket.org/galaxy/galaxy-dist#stable |
|
| upgrade: | $ hg pull $ hg update latest_2014.06.02 |
|
Tool Shed June 2, 2014 Release
A corresponding version of the ToolShed was also released on June 2, 2014. Highlights include
- Preparing for a New Main Page
- Tool Dependency Installation Recipe Enhancements
- Tool Shed API Enhancements
- Bootstrapping a New Development Tool Shed
BioBlend and CloudMan
BioBlend 0.4.3 was released on April 11, 2014.
The most recent version of CloudMan was released in January 2014.
Galaxy Community Hubs

| Share your experience now |
One new Log Board entry was added in June:
The Community Log Board and Deployment Catalog Galaxy community hubs were launched last your. If you have a Galaxy deployment, or experience you want to share then please publish them.
ToolShed Contributions
Galaxy Project ToolShed Repos
In no particular order:
Tools
-
From qfab
- pynast: PyNAST is a sequence aligner for adding new 16S rDNA sequences to existing 16S rDNA alignments - GVL
- fasttree_linux_64bit: FastTree infers approximately-maximum-likelihood phylogenetic trees from alignments of nucleotide or protein sequences - GVL
- rarefaction: Rarefaction calculation based on mothur’s rarefaction.single command - GVL
-
From crs4
- hadoop_galaxy: Hadoop-Galaxy integration
-
From evan
- bwa_wrappers: Galaxy wrappers for the BWA short read aligner.
-
From anton
- vcfprimers: Extract flanking sequences for each VCF record
- vcffixup: Count the allele frequencies across alleles present in each record in the VCF file.
- vcfsort: Sort VCF dataset by coordinate
- vcfallelicprimitives: Splits alleleic primitives (gaps or mismatches) into multiple VCF lines
- vcfaddinfo: Adds info fields from the second dataset which are not present in the first dataset.
- plus 18 more VCF related tools from anton
-
From superyuan
- refeditor: Produces a personalized diploid reference genome based on all known genetic variants of that particular individual.
-
From iracooke
- make_protein_decoys: Generate a decoy database from an input set of protein sequences. Decoys generated using this tool can be used for tandem ms searches.
- proteindb_from_gff3: Convert Augustus Generated gff3 to a Protein Database
- protxml_to_gff: Map peptides from a protXML file to genomic coordinates
- sixframe_translate: Translates sequences in a nucleotide fasta file to protein
-
From hyungrolee
- mgescan: MGEScan: Identifying long terminal repeats (LTR) and non-LTR retroelements in eukaryotic genomic sequences.
-
From iuc
- samtools_sort: Sort alignments by leftmost coordinates or read name.
-
From devteam
- bamleftalign: utility for leftaligning indels in BAM datasets. Based on bamleftalign utility for FreeBayes package.
Packages / Tool Dependency Definitions
-
From qfab
- package_numpy_1_8: Tool dependency definition; downloads and compiles the python numpy package 1.8.1 - GVL
- package_pycogent_1_5_2: Tool dependency definition; installs the PyCogent package version 1.5.2 and its dependencies - GVL
- package_uclust_1_2_22q: Tool dependency definition; installs uclust v1.2.22q for PyNAST - GVL
- package_mothur: mothur is an open-source, expandable software to fill the bioinformatics needs of the microbial ecology community
- collector_curve: Collector’s curve calculation based on mothur’s collect.single command - GVL
-
From biopython
- package_biopython_1_64: Downloads and compiles version 1.64 of the Biopython package.
-
From jankanis
- package_libxml2_2_9_1: fork of existing package_libxml2_2_9_1 from devteam with some additional environment variable exports
-
From iuc
- package_gnu_coreutils_8_22: downloads and compiles version 8.22 of the GNU coreutils.
- package_libpng_1_2: Provides the 1.2.x branch of libpng for compatibility with older software.
- package_tpp_4_6_3: Downloads and compiles version 4.6.3 of the Trans-Proteomic Pipeline.
-
From iracooke
- package_protk_1_2_6: Installs the version 1.2.6 of the protk rubygem
-
From anton
- package_vcflib: Compiled binary files for vcflib toolkit.
-
From jeremie
- package_pindel_0_2_5: downloads and compiles version 0.2.5 of Pindel.
-
From devteam
- freebayes_0_9_14_8a407cf5f4: Dependencies for FreeBayes and LeftAlign wrappers
Tool Updates:
- From vipints
- fml_gff3togtf: Uploaded version 2.0.0 of gfftools to integrate local Galaxy instances.
- From peterjc
- blastxml_to_top_descr: Uploaded v0.1.0, now also handles extended tabular BLAST output.
Other News
- The Galaxy Tool Shed: Leveraging Community Contributions with Repository Capsules
- The Galaxy Tool Shed: A Framework for Building Galaxy Tools
- Galaxy Wikipedia page, in Hebrew! Many thanks to מרים (Miriam) for creating it.
- After 8+ years & 8100+ posts the Galaxy-User mailing has retired & passed the user support torch to Galaxy Biostar
- The Galaxy Project’s public server at http://usegalaxy.org has reached 50,000 registered users! **Thank you for using Galaxy. **








