June 2013 Galaxy Update
On this page
Welcome to the June 2013 Galaxy Update, a monthly summary of what is going on in the Galaxy community. Galaxy Updates complement the Galaxy Development News Briefs which accompany new Galaxy releases and focus on Galaxy code updates.
New Public Servers
ODoSE and CBiB Galaxy joined the list of over 30 publicly accessible Galaxy servers in May.
ODoSE
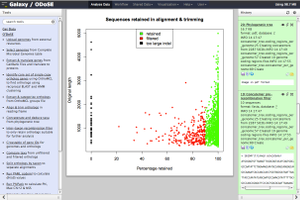
The Ortholog Direction of Selection Engine (ODoSE) supports genome-wide calculation of adaptive divergence in prokaryotes:
“The web-based graphical user-interface ODoSE (Ortholog Direction of Selection Engine) identifies all orthologs between a set of strains and allows the calculation of a novel extension of the [McDonald Kreitman] MK test, the Direction of Selection (DoS) statistic as well as the calculation of the Neutrality Index (NI) statistic for all genes shared by two taxonomic groups combined. The engine also generates the site frequency spectrum for each gene allowing one to apply more advanced methods for estimating the distribution of fitness effects and rates of adaptive evolution.”
See ODoSE: A Webserver for Genome-Wide Calculation of Adaptive Divergence in Prokaryotes by Michiel Vos, Tim A. H. te Beek, Marc A. van Driel, Martijn A. Huynen, Adam Eyre-Walker, Mark W. J. van Passel; PLoS ONE, Vol. 8, No. 5. (6 May 2013), e62447, doi:10.1371/journal.pone.0062447
CBiB Galaxy
CBiB Galaxy is a general purpose Galaxy instance that includes EMBOSS (a software analysis package for molecular biology) and fibronectin (diversity analysis of synthetic libraries of a Fibronectin domain). For now, the computational resources of the Galaxy instance of the CBiB are relatively limited. However, a migration to a dedicated server will be made soon.
CBiB Galaxy is sponsored by Centre de Bioinformatique de Bordeaux.
New Papers
39 new papers were added to the Galaxy CiteULike Group in May, including:
-
“The Genomic HyperBrowser: an analysis web server for genome-scale data” by Geir K. Sandve, Sveinung Gundersen, Morten Johansen, et al.; Nucleic Acids Research (30 April 2013)
-
“ODoSE: A Webserver for Genome-Wide Calculation of Adaptive Divergence in Prokaryotes” by Michiel Vos, Tim A. H. te Beek, Marc A. van Driel, Martijn A. Huynen, Adam Eyre-Walker, Mark W. J. van Passel; PLoS ONE, Vol. 8, No. 5. (6 May 2013), e62447
There are now almost 1000 papers in the Galaxy CiteULike Group.
Who’s Hiring
The Galaxy is expanding! Please help it grow.
- The Galaxy Project is hiring software engineers and post-docs! at both Emory and Penn State.
- Senior Developer, Stem Cell Bioinformatics Core, Sage Bionetworks, Seattle, WA, United States
- Bioinformatics Support Group Leader @ LSU
Got a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
GCC2013
| Register by 14 June & avoid late fees |
|---|
The 2013 Galaxy Community Conference (GCC2013) will be held 30 June through July 2 in Oslo Norway, at the University of Oslo.
GCC2013 is an opportunity to participate in two full days of presentations, discussions, poster sessions, keynotes, lightning talks and breakouts, all about high-throughput biology and the tools that support it. The conference also includes a Training Day for the second year in a row, this year with more in-depth topic coverage, more concurrent sessions, and more topics.
Register by 14 June and and avoid the 100% fee increase for late registration. Registration is very affordable, with combined registration (Training Day + main meeting) starting at ~ €165 for post-docs and students. Registering now also assures you a spot in the Training Day workshops you want to attend. Once a session becomes full, it will be closed to new registrations. Note that you can still register after 14 June, it will just cost you much more.
Birds of a Feather Sessions
Past Galaxy Community Conferences have been the event for networking in the Galaxy: There is no better place to meet and learn from others doing high-throughput biology. GCC2013 extends this tradition by including Birds of a Feather (BoF) meetups at the event. Birds of a Feather meetups are informal gatherings where participants group together based on common interests. If you have something in the list at right you want to meet about, or you have a whole new topic, then please join or start a GCC2013 BoF.
Sponsorships
We are pleased to announce that BioTeam is a Gold Sponsor of GCC2013.
There are still Silver and Bronze sponsorships available. Please contact the Organizers if your organization would like to help sponsor this event.
Software Carpentry Boot Camp
Directly following GCC2013, there is a unique possibility to attend a two-day Software Carpentry Boot Camp at the University of Oslo (in a building close to where the GCC is held). Software Carpentry Boot Camps aim to to help scientists and engineers become more productive by teaching them basic computing skills like program design, version control, testing, and task automation. In this two-day boot camp, short tutorials will alternate with hands-on practical exercises.
The course is aimed at postgraduate students and other scientists who are familiar with basic programming concepts (like loops, conditionals, arrays, and functions) but need help to translate this knowledge into practical tools to help them work more productively.
Content: The syllabus for this boot camp will include:
- using the shell to do more in less time
- using version control to manage and share information
- basic Python programming
- how (and how much) to test programs
Visit the Boot Camp Page for more information, and registration.
Other Upcoming Events
See the [Galaxy Events Google Calendar](http://bit.ly/gxycal) for details on these and other events.Events
GalaxyAdmins
The May GalaxyAdmins Meetup featured Andrew Warren of the Virginia Bioinformatics Institute at Virginia Tech speaking on their Galaxy deployment RNA Rocket for Pathogen Portal. Dannon Bakeralso gave a Galaxy Project update. Slides and a screencast are now available.
Duplicate Accounts on Main
The terms of use for the Galaxy Project’s free Galaxy server, http://usegalaxy.org (a.k.a. Main) specify
In the past we have dealt with multiple accounts on an ad-hoc basis. In May 2013, an automated system was put into place to detect, disable, and eventually delete duplicate accounts. If the automated system determines that an account is a duplicate, the account will be disabled and the account owner will receive an email asking if this is the account they wish to keep, and if so, to follow the directions in the email.
If your account is suspended and you believe that you are not in violation of Main’s terms of use, then please contact [terms_and_conditions AT galaxyproject DOT org](mailto:terms_and_conditions AT galaxyproject DOT org).
Galaxy Distributions
The most recent official distribution was on April 1, 2013. However, a security update was released on April 8. See this news item for details. We are planning the next release for early June.
Tool Shed Contributions
- rxlr_venn_workflow: a workflow for comparing 3 RXLR prediction methods with a Venn Diagram
- vcf_to_variantdb: Send UnifiedGenotyper VCF to a !VariantDB instance
- iquant: isobaric tag quantification program
- appendfdr: compute false discovery rate of tabular data using decoy entries
- bumbershoot: Tabb lab tools idqonvert, myrimatch, tagrecon, all for protein identification.
- dbbuilder: download protein databases from common sources
- nbic_fasta: Galaxy-P unofficial fork of NBIC Galaxy tools for FASTA manipulation
- maxquant: large mass-spec data quantitative proteomics analysis, especially high-resolution mass spectometry
- msconvert: Tool wrappers for msconvert, distributed as part of Proteowizard.
- peptideshaker: Identify proteins combining X! Tandem, OMSSA (using SearchGUI) and PeptideShaker
- psm_eval: Peptide-Spectrum-Match reevaluation tool
- allele_counts: Parse Naive Variant Detector output, count alleles and calculate minor allele frequencies.
- snp_analysis_conversion: Format conversion for SNP Analysis tools from Miller Lab @ PSU.
Other News
- Just discovered these tutorials by Lance Parsons on RNA-Seq, ChIP-Seq, SNP & Indels, and IGV all using Galaxy
- Scaling variant detection pipelines for whole genome sequencing analysis
- There is now a Google+ Galaxy Group. Thanks to Samuel Lampa for setting this up.