On this page
In the November 2020 issue
- JXTX: The James P. Taylor Foundation for Open Science
- Event news
- Galaxy platform news: GalaxyTrakr, Galaxy Africa, Australia, Europe, and UseGalaxy.*
- Blog posts: Advanced Microbiology with Galaxy and TIaaS
- Training material and doc updates
- Publications
- Q: Who’s hiring? A: Eight different groups
- New releases
- Other news too
If you have anything to include to next month’s newsletter, then please send it to outreach@galaxyproject.org.
JXTX: The James P. Taylor Foundation for Open Science

The foundation continuing James’s legacy of supporting Open Science has been renamed from JTech to JXTX: The James P. Taylor Foundation for Open Science. The JXTX Foundation will continue to focus on mentoring and enabling junior researchers to participate in the Open Science James was dedicated to.

The JXTX Foundation’s first activity is to sponsor 10 graduate students to attend the 2020 Biological Data Science Conference at Cold Spring Harbor Laboratory. Awards were competitive and we are delighted to support this first cohort of awardees.

You can help continue James’s legacy of open and reproducible science by contributing to JXTX: The James P. Taylor Foundation for Open Science.
Event News
Despite COVID-19, there is still a lot going on. Some of it is virtual, but live events are starting to happen again, especially in Europe. We have updated our list of events to reflect what we know. Some highlights:

There will be one roundtable meetup this month, on November 12. Discussion will focus on working with the Galaxy Training Network.

Paper Cuts are annoying but easy to fix bugs. The first Paper Cuts event in October was a smashing success, so we are going to do it again on November 18. Our second one-day Paper Cuts contribution fest will also be a 24-hour event spanning all time zones with our worldwide community.
Please save the date! It’s an ideal opportunity for newcomers to become a Galaxy contributor.

This November 10 webinar by Barbara Arbeithuber and Nick Stoler will present Du Novo Sequencing, a method that can achieve <1% variant detection, by removing the reliance on a reference sequence, preserving a higher proportion of the input reads, and being available for Galaxy.

Three Galaxy Australia events are being offered by in November:
- Webinar: Galaxy Australia: enabling online data analysis for the research community, 4 Nov, from Australian BioCommons.
- Workshop: Online data analysis for biologists, 11 Nov, from Australian BioCommons.
- Workshop: Hybrid de novo genome assembly - Nanopore and Illumina, 30 Nov, from Melbourne Bioinformatics.

19th of November, Online
A dedicated day of collaborative work on the training content and calls with the Galaxy training community. Anyone who would like to contribute, learn how to contribute to the Galaxy Training Material or just catch up with the community is very welcome to join.

3rd of December, Online
This hands-on workshop will familiarize you with the Galaxy user interface & execute (label-free) proteomics data-analysis. Taught by Melanie Föll and Matthias Fahrner from the University of Freiburg.

7 au 10 décembre 2020, en visio
Un Hackathon pour partager des compétences en terme du développement logiciel et d’administration système des Interactive Tools de Galaxy. Limité à 25 participants.

Galaxy Admin Training is coming in January. It will be online and global, and registration will open this month. Watch Twitter, this web site, and Galaxy Matrix channels for the announcement.

There are more than 16 events before the end of the year:
- workshops, CoFests, talks, …
- all of them are online.
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. Here’s the recent platform news we know about:

The GalaxyTrakr server is provided by the US Food and Drug Administration to support food-borne pathogen research worldwide. A User Guide, FAQ, Videos, and Email support are available. An account is required and anyone working in public health can apply.

Devoted to assisting researchers on the African continent, accelerate their genomics research and analyses. This public server is configured by the Galaxy Africa team, and hosted by the UseGalaxy.eu team.

Galaxy Australia is now officially supporting 10,000 researchers to analyse their valuable data. It’s rapidly becoming the essential research tool for so many around the world.

Galaxy EU has surpassed 20,000 users!
5 continents, 100 countries, more than 100 industry companies, 4,000 users trained using TIaaS, 11 million jobs, 12 jobs per minute…
UseGalaxy.* News

- Talking about UseGalaxy.eu in a blog post or presentation? Galaxy Europe has provided cool graphics and a fact sheet as well.
- Lots of tool updates on UseGalaxy.eu and UseGalaxy.org.au.
Galactic Blog Activity
By Leighton Pritchard.
Leighton reports on the University of Strathclyde’s experience teach Advanced Microbiology using UseGalaxy.eu’s Training Infrastructure as a Service (TIaaS).
Spoiler: It went well.

Doc, Hub, and Training Updates
By Subina Mehta, Timothy J. Griffin, Pratik Jagtap, Emma Leith, Marie Crane, and Praveen Kumar
A three part series on doing metaproteomics analysis with metaQuantome:
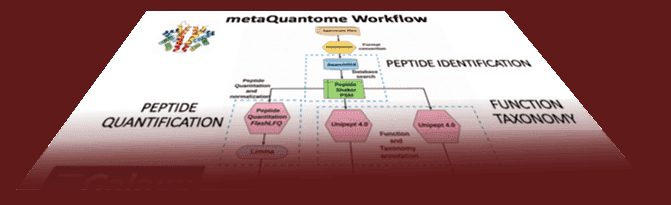
Based on the work by Delphine Larivière and James Taylor with their COVID-19 Lectures, we have implemented a similar feature in the Galaxy Training Network. Learn how to use it.

This tutorial got a thorough update from Mallory Freeberg.

Publications
Pub curation activities are on hiatus right now but a few publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library anyway. Here are the new open access Galactic and Stellar pubs:
Rivière, E., Heupink, T. H., Ismail, N., Dippenaar, A., Clarke, C., Abebe, G., Heusden, P., Warren, R., Meehan, C. J., & Van Rie, A. (2020). Briefings in Bioinformatics. doi: doi.org/10.1093/bib/bbaa246
VijayKrishna, N., Joshi, J., Coraor, N., Hillman-Jackson, J., Bouvier, D., Beek, M. van den, Eguinoa, I., Coppens, F., Golitsynskiy, S., Stolarczyk, M., Sheffield, N. C., Gladman, S., Cuccuru, G., Grüning, B., Soranzo, N., Rasche, H., Langhorst, B. W., Bernt, M., Fornika, D., … Blankenberg, D. (2020). BioRxiv, 2020.10.09.327114. doi: 10.1101/2020.10.09.327114
Tekman, M., Batut, B., Ostrovsky, A., Antoniewski, C., Clements, D., Ramirez, F., Etherington, G. J., Hotz, H.-R., Scholtalbers, J., Manning, J. R., Bellenger, L., Doyle, M. A. , Heydarian, M., Huang, N., Soranzo, N., Moreno, P., Mautner, S., Papatheodorou, I., Nekrutenko, A., Taylor, J., Blankenberg, D., Backofen, R., Grüning, B. (2020). GigaScience, 9(10). doi: 10.1093/gigascience/giaa102
Timme, R., Laxmi, S., Balkey, M., Randolf, R., Wolfgang, W., & Strain, E. (2020). Protocols.io. doi: 10.17504/protocols.io.bdvfi63n
de Koning, W., Miladi, M., Hiltemann, S., Heikema, A., Hays, J. P., Flemming, S., van den Beek, M., Mustafa, D. A., Backofen, R., Grüning, B., & Stubbs, A. P. (2020). GigaScience, 9(10). doi: 10.1093/gigascience/giaa105
Goonasekera, N., Mahmoud, A., Chilton, J., Afgan, E. (2020). Bioinformatics. doi: 10.1093/bioinformatics/btaa860
Who’s Hiring
Università degli Studi di Bari Aldo Moro, Regione Puglia, Italy.
Development and implementation of a computing platform in a cloud environment with a web interface based on the Galaxy workflow management system to provide access to computational tools for Precision Medicine compliant with GDPR directives.
Applications close 14 November.

Roche, Bay Area, California, United States.
- Lead data mining for biomarker discovery for medical conditions of interest.
- Develop Agile Assay Design (AAD) tools for qPCR tests.
- NGS data analysis tools and/or workflows.
- Use these tools & workflows for R&D projects.
- Deploy these tools on Roche intranet (Galaxy) and train scientists to use them.

ELIXIR Norway Team, University of Oslo, Norway.
Join the ELIXIR development / operations team at the University of Oslo (UiO), participating in the development, implementation and operation of functionality for various national and international solutions provided by ELIXIR Norway, including UseGalaxy.no and NeLS.

Johns Hopkins University, Baltimore, Maryland, United States.
Provide technical expertise and oversight for the AnVIL Project, which incorporates Galaxy, Bioconductor, Terra, Gen3, and Dockstore into a secure cloud-based software ecosystem for genomic data analysis.

Help with Galaxy-P tools and workflow development, and general support for Galaxy implementation at the Minnesota Supercomputing Institute.

The Blankenberg Lab in the Genomic Medicine Institute at the Cleveland Clinic Lerner Research Institute is searching for Software Engineers and Postdoctoral Fellows. We utilize high-throughput omics technologies, such as Next-Generation Sequencing, and data-intensive computing to explore biomedical research questions.

A 3-year position of genomic data analyst is available to work within the “COllaborative NEtwork on research for Children and Teenagers with Acute Myeloblastic Leukemia” CONECT-AML framework.
The project is funded by the Institut National de Recherche sur le Cancer (INCA) and the Fight Kids Cancer programme. It involves 10 participant teams across France, with clinical or fundamental approaches.

The Schatz Lab at Johns Hopkins University is looking for:
- Self-driven individuals that can work independently to fill multiple software development positions on the Galaxy Project.
- Ambitious individuals to fill a programmer analyst position working on the Galaxy and AnVIL projects.

Releases
Galaxy Language Server implements the Language Server Protocol to assist in the development of Galaxy tool wrappers inside modern code editors.
Please note this is still an early work in progress and bugs and issues are expected.
It provides realtime XML validation, code completion, help/documentation and other smart features to help in following best practices during the development process of XML tool wrappers for Galaxy.
The Galaxy Tools Visual Studio Code extension that uses the Galaxy Language Server can be downloaded from the Open VSX Registry and the Visual Studio Marketplace.
Other news
A group of Freiburg researchers provide global, open access to data on the SARS-CoV-2 genome which could hold the key for a new approach to treating the virus.




