On this page
Hello all,
October brings Galaxy involvement in Hacktoberfest, and Outreachy, plus a nice batch of trainings, talksm and a Galaxy Papercuts CoFest day too. It also brings news of new job openings on two continents, two new platforms, blog posts (where My Little Pony makes an appearance, really), GTN updates (few in number but big in their coolness), publications, and releases.
In other words, another active month in the Galaxy… :-)
Beatriz Serrano-Solano and Dave Clements, Editors
PS: Subscribe to the Galaxy Announce mailing list and receive an email whenever this newsletter is published.
Community News

9-10 November
This 2 day pre-Inbix2021 workshop will feature talks and hands-on exercises, and the formal launch of the Galaxy India Community.
Presenters include include Christopher B Barnett, Anshu Bhardwaj, Simon Bray, Sona Charles, Dave Clements, Stephan Flemming, Selvaraman Nagamani, and Gareth Price.
Early registration ends November 5. Bioclues members can register for free.
Hacktoberfest is an annual global contribution drive for open source software, and is underway now. Sign up and make a pull request for documentation, new tools, or contribute to any of the Galaxy GitHub repositories to earn a free Hacktoberfest T-shirt.
See the announcement for details.
Galaxy will be participating in the December 2021 Outreachy Internship Cohort. Outreachy is an internship program for open source communities and non-profits. Galaxy has two proposed projects for this cohort.
Interested in helping? See the announcement for details on what you can do, starting right now.

Two new positions were listed in the Galaxy Career Center recently. The Career Center lists any open Galaxy related positions that we know about. Currently there are openings at
- Helmholtz Zentrum München, Germany
- Tufts University in Massachusetts
- Sciensano in Belgium
- NEB in Massachusetts, US
- GalaxyWorks in the US
- Cleveland Clinic in Ohio, US
- Johns Hopkins University in Maryland, US
Event News
See all upcoming events here:
20 October, Online
A workshop to build capacity in SARS-CoV-2 data analysis! It will be a 1-day event with pre-recorded lectures, hands-on and demos. During the workshop, there will be live support in chat and live Questions & Answers sessions.
Registration will open soon, stay tuned!

19-21 October, Bordeaux, France
- Maîtriser les outils d’analyse de séquences tels que les alignements et savoir interpréter les résultats
- Maîtriser les formats et les analyses des nouvelles données issues du séquençage (NGS)
- Savoir utiliser l’environnement Galaxy

The Galaxy Developer Round Table meetup in October is:
October 28: Image analysis in Galaxy - pain points and lessons learnt led by Yi Sun and Jean-Karim Hériché of EMBL.
We are seeking volunteers (like you!) to lead the discussion on your favorite topic at future roundtables.

21 October, Online, Global
Please join us for the CoFest day on October 21 to help the Galaxy Ecosystem become a better place, and to help new contributors come on board.
This month the Metabolomics community will meet to work on tools, get in touch with Melanie Föll to participate.
We will be on Matrix for chat all day long, please take advantage of both to communicate with your collaborators around the world.

15-24 November, Online, India. Apply by 17 October.
Microbiome experts and instructors from the Galaxy community will lead this online course on microbiome analysis via interactive resources. Presenters include Sanjeev Khosla, Anshu Bhardwaj, Manoj Kumar, Robert Hettich, Pratik Jagtap, Dave Clements, Subina Mehta, Saskia Hiltemann, and many, many more.

29-30 November, Online, UK.
Curso de análisis de células únicas con Galaxy en español. ¡Pronto se abrirá el periodo de inscripción!
—
Single-cell workshop in Spanish. Registration will open soon!
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. Here’s the recent platform news we know about:
MitoLink is an integrated workflow system to facilitate understanding of genotype-phenotype correlations in cases of mitochondrial dysfunction. MitoLink provides a complete pipeline based system to analyse and visualize the role of SNPs in Mitochondrial Diseases. It also provides a MitoLink User Manual, shared data, and workflows.
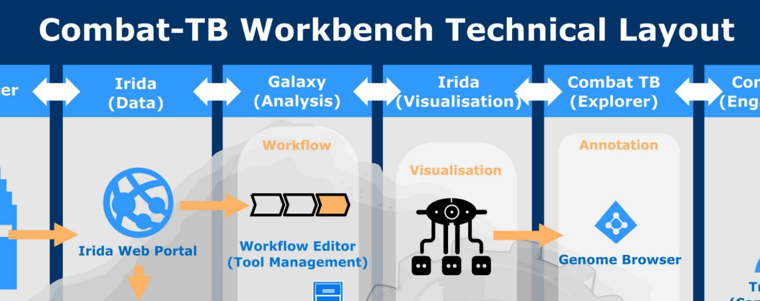
COMBAT-TB is a modular, easy to install application that provides a web based environment for Mycobacterium tuberculosis bioinformatics. It is built using two main software components: the IRIDA Platform for its web-based user interface and data management capabilities and the Galaxy bioinformatics workflow platform for workflow execution.
VEuPathDB Update
The VEuPathDB Galaxy instance has been updated and now offers improved job and workflow performance for users. Additional tool such as those for scRNA-seq (Scanpy, Seurat), species ID (CryptoGenotyper), table manipulation (Datamash), and proteomics (MSstats, Search GUI, Peptide Shaker, MaxQuant) were added, and existing tools upgraded too. Give it a try.
UseGalaxy.* News

- The FTP server on UseGalaxy.eu has been upgraded, providing more space and accepting only encrypted connections (TLS).
- UseGalaxy.eu as a cross-disciplinary platform for European researchers
- Lots of tool updates on UseGalaxy.eu and UseGalaxy.org.au.
Galactic Blog Posts
By Tomas Klingström.
Tomas has shared with us his hilarious experience when running a workshop from home. Find out the whole story!

By Enis Afgan.
The Human Genetics working group focuses on expanding Galaxy’s ability to analyze protected human data. Currently, this is made possible via AnVIL where Galaxy is deployed in a FedRAMP certified environment and alongside 3PB of data. This is the only such environment in the world where anyone can sign up and start working with this data within minutes.
Doc, Hub, and Training Updates
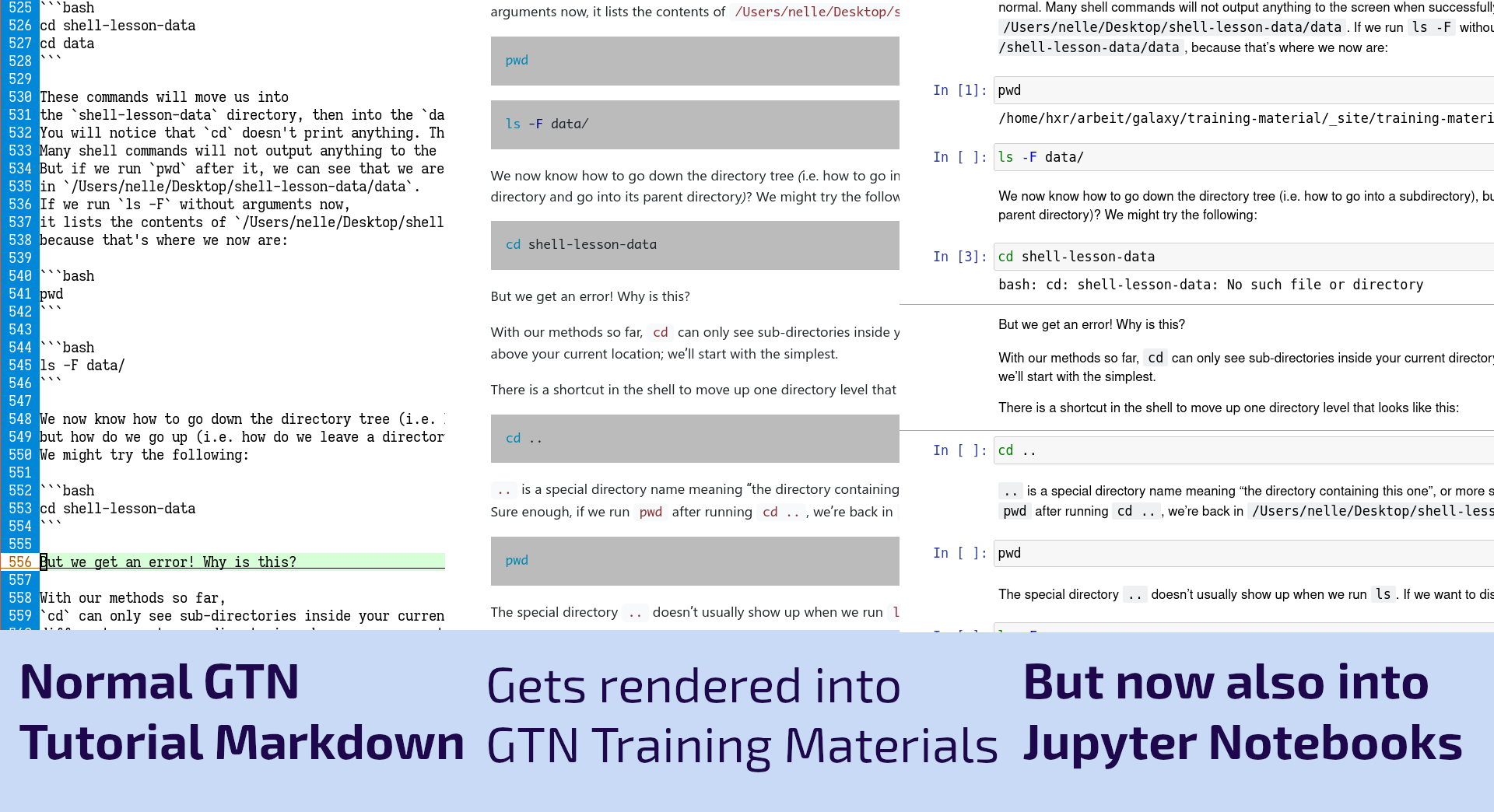
Check out the first GTN tutorial to use Jupyter notebooks for coding and the command line, and that themselves run inside Galaxy. (It’s OK, you can read that sentence again.) Using Jupyter notebooks greatly reduces the cutting and pasting in these. See the full announcement for details and link to documentation on how to use this approach.
The GTN library has a new section, the Data Science Survival Kit for tutorials focused on Data Science. Initially, it has three tutorials in it:
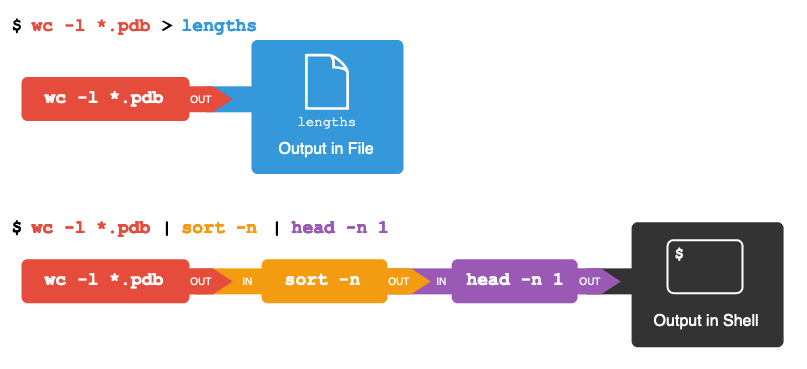
- CLI basics in Galaxy , by The Carpentries, Helena Rasche, and Bazante Sanders.
- Advanced CLI in Galaxy , by The Carpentries, Helena Rasche, and Bazante Sanders.
- CLI Educational Game - Bashcrawl, by Helena Rasche.
This introductory tutorial got a major update from Cristóbal Gallardo. Screenshots were updated, and many explanations were clarified.
The Galaxy Training Network would like to get feedback from instructors on the use of Galaxy for training, the preparation needed and the Galaxy Training Material.
Publications
Pub curation activities are on a semi-hiatus right now but a few publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library anyway. Here are the new open access Galactic and Stellar pubs:
Föll, M. C., Volkmann, V., Enderle-Ammour, K., Wilhelm, K., Guo, D., Vitek, O., Bronsert, P., & Schilling, O. (2021). BioRxiv, 2021.08.09.455649. https://doi.org/10.1101/2021.08.09.455649
Maier, W., Bray, S., van den Beek, M. et al. Ready-to-use public infrastructure for global SARS-CoV-2 monitoring. Nat Biotechnol 39, 1178–1179 (2021). https://doi.org/10.1038/s41587-021-01069-1
Heusden, P. van, Mashologu, Z., Lose, T., Warren, R., & Christoffels, A. (2021). MedRxiv, 2021.09.23.21263983. https://doi.org/10.1101/2021.09.23.21263983
Releases

Bug fixes and the documentation has been improved in the latest version of Planemo.
- Fix documentation to include
--download_outputsflag (thanks to@simonbray).Pull Request 1184 - Select refgenie config based on Galaxy version
Pull Request 1187_ - Extend autoupdate subcommand to workflows (thanks to
@simonbray).Pull Request 1151
This release contains the Galaxy Language Server and the Galaxy Tools Visual Studio Code Extension.
The standalone version of the language server is available as a PyPI package.
The Galaxy Tools Extension is available at Open VSX Registry and Visual Studio Marketplace.
Other News
ELIXIR Germany has published the handout “ELIXIR GERMANY - Shaping the Bioinformatics Infrastructure for Life Sciences in Europe”.