June 2019 Galactic News
GCC2019 registration / Poster/Demo abstracts / sponsors, GTN CoFest, Platforms, Pubs, Jobs, Blog!, Training, Tools, Releases and more!
On this page
The June 2019 Galactic News is here! This is a summary of what is going on in the Galaxy community.
- GCC2019:
- Plus 13 other upcoming events in the next 90 days
- 150 new publications, great resources lead to great insight.
- Galaxy Platform News: New platforms for tranSMART, antibiotic resistance, phylogeny, long-read data, mass spectrometry Imaging, and Rice!
- A new entry to The Galactic Blog on Galaxy cloud bursting
- At least 8 Open positions in five countries on two continents.
- Updates to training materials and documentation.
- New tools and new releases.
- And some cool other news too:
- The new Galaxy Services Status Web Site
- Black Duck Open Hub updates it’s Galaxy project stats and we are doing great.
If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
GCC2019, 1-8 July, Freiburg, Germany
GCC2019 will be held the first week of July in Freiburg, Germany. The tenth GCC features more presentations and more training than any GCC before it. If you are working in data intensive life science research then there will not be a better place to share your work, learn from others, and find new collaborators.
Advance Registration ends 7 June - this Friday
Advance registration for the 2019 Galaxy Community Conference (GCC2019) is ends this this Friday, 7 June. Register this week and avoid late registration fee hikes that start on Saturday morning. (Insert your favorite Saturday morning hangover joke here.)
Poster & Demo Abstract Submission Deadline: 10 June
Abstracts for poster presentations and software demonstrations are still being accepted for consideration. Got something to show? Submit your abstract before the 10 June deadline!.
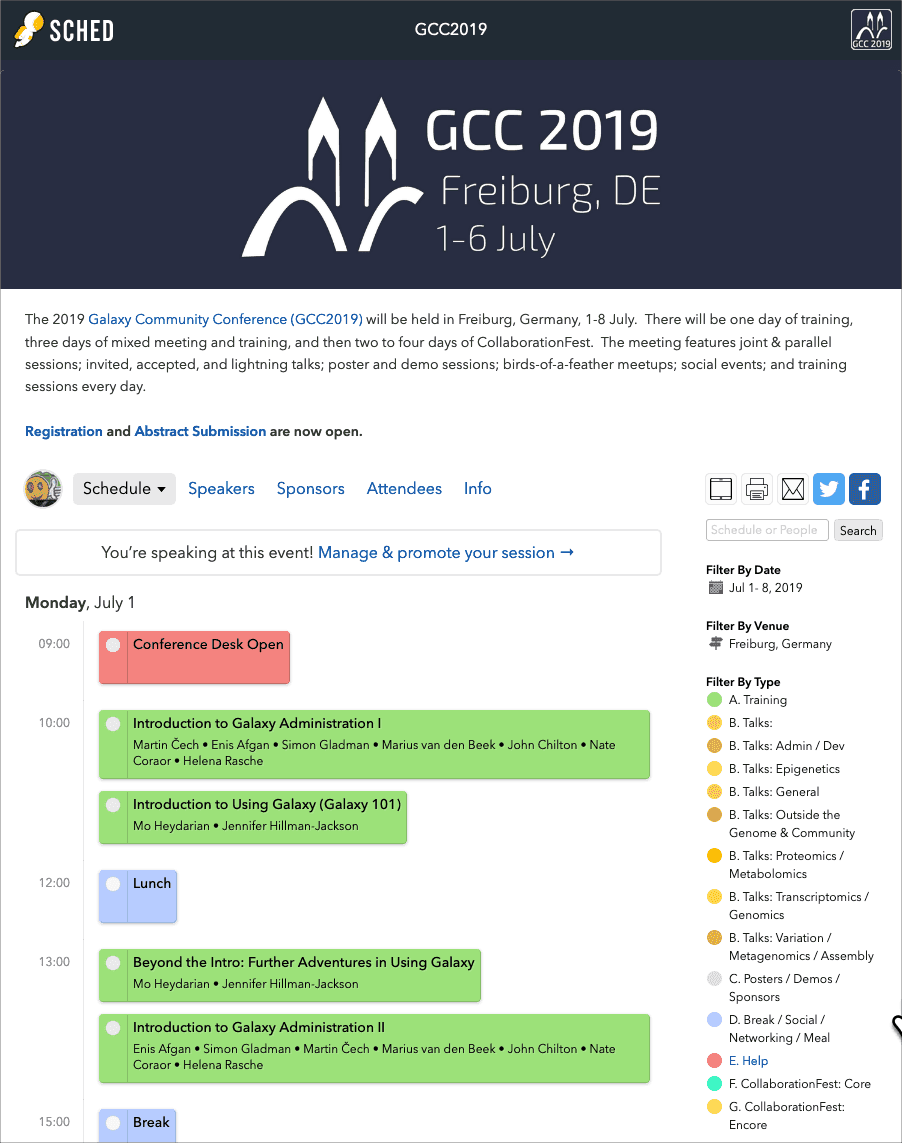
Conference Schedule is Online
The full conference schedule is now online. There are
- Over 35 training topics ranging from Beyond the Intro: Further Adventures in Using Galaxy to Visualisation development in Galaxy
- 15 Sessions, including dedicated sessions on
- Epigenetics, Galaxy Administration & Development (3 sessions), Outside the Genome & Community (3 sessions), Proteomics and Metabolomics, Transcriptomics & Genomics (2 sessions), Variation / Metagenomics / Assembly
- Five invited talks:
- Evolution of X chromosome recognition during Drosophila Dosage Compensation, Claudia Isabelle Keller Valsecchi, Max Planck Institute of Immunobiology and Epigenetics
- Pou5f3, SoxB1 and Nanog control the Zygotic Genome Activation in Zebrafish by Chromatin Remodeling, Marina Veil, Albert Ludwigs University of Freiburg
- Galaxy Community Update, Jeremy Goecks, OHSU; Daniel Blankenberg, Cleveland Clinic; Anton Nekrutenko, Penn State University; James Taylor, Johns Hopkins University
- Data visualisation by citizen science participants: the case of birds and bats monitoring schemes and Galaxy-E, Romain Lorrilière, French National Museum of Natural History
- UseGalaxy: Elucidating chromatin biology of mouse cardiomyocytes, Stephan Nothjunge, University of Freiburg
- Over 70 talks
- Three Poster, Demo, and Sponsor Sessions (Posters & demos to be posted next week)
- Three hour long sessions for BoFs (call for BoFs going out next week)
- 4 days of CollaborationFest
Upcoming events
These and other Galaxy related events are coming up:
Publications
150 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month.
Highlighted Publications
There were 10 Galactic and Stellar publications added, and 9 of those are open access:
- Accessible and reproducible mass spectrometry imaging data analysis in Galaxy, Melanie Christine Föll, Lennart Moritz, Thomas Wollmann, Maren Nicole Stillger, Niklas Vockert, Martin Werner, Peter Bronsert, Karl Rohr, Björn Andreas Grüning, Oliver Schilling. bioRxiv 628719; doi: 10.1101/628719
- G-OnRamp: a Galaxy-based platform for collaborative annotation of eukaryotic genomes, Yating Liu, Luke Sargent, Wilson Leung, Sarah C. R. Elgin and Jeremy Goecks. Bioinformatics, btz309, doi: 10.1093/bioinformatics/btz309
- MetaDEGalaxy: Galaxy workflow for differential abundance analysis of 16s metagenomic data, Mike W.C. Thang, Xin-Yi Chua, Gareth Price, Dominique Gorse, Matt A. Field. F1000Research 2019, 8:726, doi: 10.12688/f1000research.18866.1)
- NGPhylogeny.fr: new generation phylogenetic services for non-specialists, Frédéric Lemoine, Damien Correia, Vincent Lefort, Olivia Doppelt-Azeroual, Fabien Mareuil, Sarah Cohen-Boulakia, Olivier Gascuel. Nucleic Acids Research, gkz303, doi: 10.1093/nar/gkz303
- Rice Galaxy: an open resource for plant science, Venice Juanillas, Alexis Dereeper, Nicolas Beaume, Gaetan Droc, Joshua Dizon, John Robert Mendoza, Jon Peter Perdon, Locedie Mansueto, Lindsay Triplett, Jillian Lang, Gabriel Zhou, Kunalan Ratharanjan, Beth Plale, Jason Haga, Jan E. Leach, Manuel Ruiz, Michael Thomson, Nickolai Alexandrov, Pierre Larmande, Tobias Kretzschmar and Ramil P. Mauleon. GigaScience, 8, 2019, 1–14, doi: 10.1093/gigascience/giz028
- The RNA workbench 2.0: next generation RNA data analysis, Jorg Fallmann, Pavankumar Videm, Andrea Bagnacani, Bérénice Batut, Maria A. Doyle, Tomas Klingstrom, Florian Eggenhofer, Peter F. Stadler, Rolf Backofen and Björn Grüning. Nucleic Acids Research, gkz353, doi: 10.1093/nar/gkz353
- Galaxy External Display Applications: Closing a dataflow interoperability loop, Daniel Blankenberg, John Chilton, and Nate Coraor. bioRxiv 642280; doi: 10.1101/642280
- Prediction and Characterization of miRNA/Target Pairs in Non‐Model Plants Using RNA‐seq, Kira C. M. Neller, Alexander Klenov, Katalin A. Hudak. Current Protocols in Plant Biology, e20090. doi: 10.1002/cppb.20090
- The Repetitive DNA Composition in the Natural Pesticide Producer Tanacetum cinerariifolium: Interindividual Variation of Subtelomeric Tandem Repeats, Jelena Mlinarec, Ana Skuhala, Adela Jurković, Nenad Malenica, Jamie McCann, Hanna Weiss-Schneeweiss, Borut Bohanec and Višnja Besendorfer. Frontiers in Plant Science, 16 May 2019, doi: 10.3389/fpls.2019.00613
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 95 | +Methods | 43 | +UsePublic | 23 | +Workbench | 18 | +RefPublic | |||
| 17 | +UseMain | 11 | +UseLocal | 9 | +IsGalaxy | 7 | +Tools | |||
| 4 | +Reproducibility | 3 | +Shared | 2 | +Education | 2 | +Project | |||
| 2 | +Visualization | 1 | +HowTo |
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. A lot was added in March:
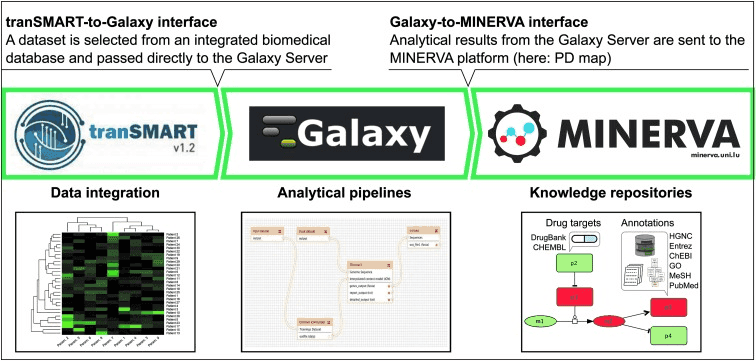
The TGM pipeline
The TGM pipeline is a platform combining tranSMART server, Galaxy Server, and MINERVA platform to enable visually-aided exploration, analysis, and interpretation of high-throughput translational medicine data. The TGM pipeline enables establishing workflows with Galaxy for data processing using tools and resources of the tranSMART server.
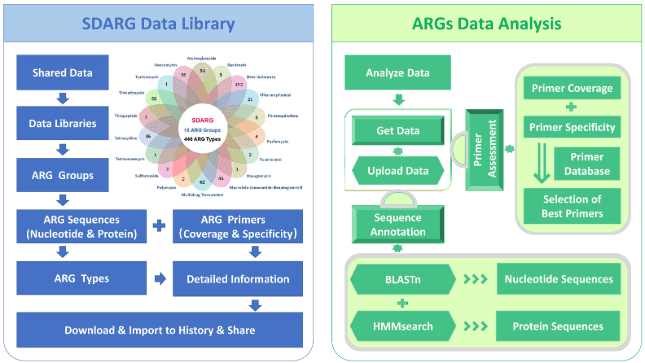
ARGA
The Antibiotic Resistance Gene Analyzer (ARGA) is a pipeline used for identification of antibiotic resistance genes using the Sequence Database for ARGS (SDARGS). The ARGA Galaxy instance provides all of the nucleotide and protein sequences in SDARG and ARGs primer databases in the Shared Data and Data Libraries sections for users. A guide for recommended usage of ARGA can be found here.
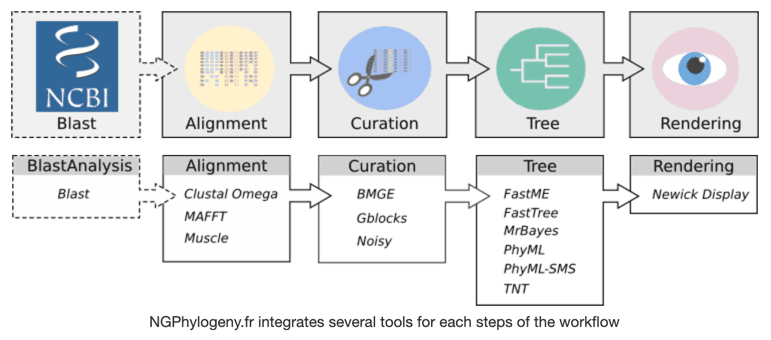
NGPhylogeny.fr
NGPhylogeny.fr is a free and simple to use web service dedicated to reconstructing and analysing phylogenetic relationships between molecular sequences. The NGPhylogeny service allows users to execute and customize Galaxy workflows while employing a novel interface to Galaxy. Additional documentation can be found here.
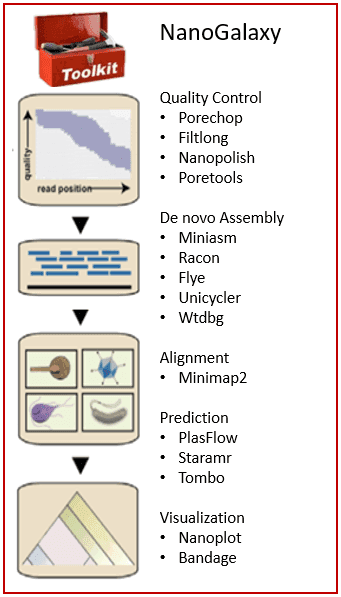
NanoGalaxy
NanoGalaxy is a webserver for processing, analysis, and visualization of long-read nucleic acid sequencing data from Oxford Nanopore Technologies (ONT) and Pacific Biosystems (PacBio) with Galaxy. This specialized Galaxy instance provides access to a collection of best practice workflows and commonly used tools for analysis of long-read sequence data.
Galaxy Workbench for Mass Spectrometry Imaging
This is a custom docker image for doing mass spectrometry image processing in a Galaxy platform. The tools available in this image are also available on the UseGalaxy.eu platform.
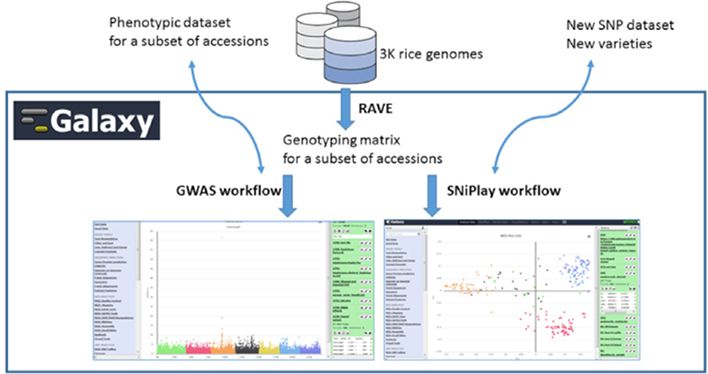
Rice Galaxy
The Rice Galaxy server is a revamped version of IRRI’s previously existing Galaxy server. It features tools for designing single-nucleotide polymorphism assays, analyzing genome-wide association studies, population diversity, rice−bacterial pathogen diagnostics, and a suite of published genomic prediction methods and has shared datasets that include high-density genotypes from the 3,000 Rice Genomes project and sequences with corresponding annotations from 9 published rice genomes.
Galaxy Platforms in Publications
We tag papers that use, mention, implement or extend public Galaxy platforms (servers, services, clouds, containers…). Here are the platforms that were referenced at least twice in the past month’s publications:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 21 | >Huttenhower | 9 | >CPT | 7 | >Workflow4Metabolomics | 5 | >RepeatExplorer | |||
| 3 | >ARGs-OAP | 3 | >Galaxy-P | 3 | >UseGalaxy.org.au | 2 | >deepTools | |||
| 2 | [>Globus Genomics](https://www.zotero.org/groups/1732893/galaxy/tags/>Globus Genomics) | 2 | >IRProfiler | 2 | >RiboGalaxy |
New Galactic Blog Post: Enabling Cloud Bursting for Galaxy
For the second month in a row we have a new Galactic Blog post from Enis Afgan and Vahid Jalili:
Who’s Hiring

The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back!
- Bioinformaticien h/f - INNOLEA, Mondonville, France. “Le développement d’outils pour l’analyse de données génétiques, génomiques et la mise en œuvre dans une plateforme Galaxy”
- Postdoctoral Fellows, Blankenberg Lab, Cleveland Clinic, Cleveland, Ohio, United States.
- Scientist - Molecular R&D, MilliporeSigma, Rockville, Maryland, United States
- ELIXIR Belgium has three Galaxy-related openings in Ghent:
- ELIXIR Open Science Community Manager, VIB-UGent Center for Plant Systems Biology
- ELIXIR Software developer data management tools, VIB-UGent Center for Plant Systems Biology
- ELIXIR Scientific Cloud Coordinator, VIB-UGent Center for Plant Systems Biology
- Software Developer, Harvard T.H. Chan School of Public Health, Boston, Massachusetts, United States. “Basic automated analysis workflows using Galaxy for 16S marker gene, metagenomic, and metatranscriptomic data leveraging existing software.”
- The The European Galaxy Team has open positions, Freiburg, Germany
- Software engineer, system analysts/administrators, data analyst, and a community and/or research manager
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Doc, Hub, and Training Updates
Updates from the Galaxy Training Materials:
- A short version of the “16S Microbial Analysis with mothur” was added by Saskia Hiltemann. This complements the longer version. You can switch at any point from the extended one to the short one. Which is great if you are running out of time.
- Three new tutorials were added by Chris Barnett, Tharindu Senapathi and Simon Bray, providing an introduction to performing molecular dynamics simulations and analysis in Galaxy.
- Tired of individually loading training datasets from Zenodo? Tutorials can now support bulk copying of URL lists. Thanks to Helena Rasche.
Hub updates:
- Data Privacy in Galaxy: A new page describing how to use Galaxy’s data privacy features; from Martin Cech
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)Tool Shed contributions in April and May 2019.
Releases
New additions to (and editions in) the Galaxy Ecosystem.
Planemo 0.60.0
Planemo is a set of command-line utilities to assist in building tools for the Galaxy project. These releases included numerous fixes and enhancements.
See GitHub for details.
Pulsar 0.11.0
Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
ephemeris 0.10.0
Ephemeris is a small Python library and set of scripts for managing the bootstrapping of Galaxy plugins - tools, index data, and workflows. It has extensive documentation. This release features numerous newly contributed features.
Other News
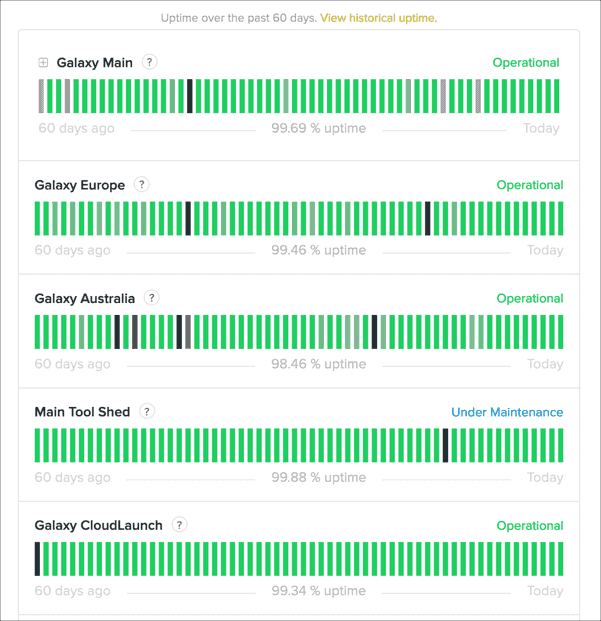
Galaxy Services Status Web Site
Automated status tracking of our most important services is available at status.galaxyproject.org. This is updated in real time, and will be used to post status updates whenever there are issues.
Black Duck Open Hub updates it’s Galaxy project stats
Black Duck’s Open Hub web site reports stats on open source projects such as Galaxy. They just updated their summary of Galaxy:
- Over the past twelve months, 198 developers contributed new code to Galaxy Bioinformatics Platform. This is one of the largest open-source teams in the world, and is in the top 2% of all project teams on Black Duck’s Open Hub.
data.galaxyproject.org directory view.
From Enis Afgan:
- If you ever wondered what reference data is available on the Galaxy Project CVMFS at data.galaxyproject.org, here’s a snapshot file system directory view, and here’s the equivalent navigable view.