May 2019 Galactic News
GCC2019 registration / Poster/Demo abstracts / sponsors, GTN CoFest, Platforms, Pubs, Jobs, Blog!, Training, Tools, Releases and more!
On this page
The May 2019 Galactic News is here! This is a summary of what is going on in the Galaxy community.
- Events
- GCC2019:
- Early registration ends 17 May
- Poster and Demo Abstract Submission is open
- Travel Fellowships: 2nd round ends this Friday
- GigaScience returns as a GCC sponsor!
- GTN CoFest and Community Call: 16 May
- Plus 17 other upcoming events in the next 90 days
- GCC2019:
- 265 new publications, great resources lead to great insight.
- Some most excellent Galaxy Platform News, including a single cell ‘omics workbench, network analysis, subcellular spatial organization modeling, and more!
- A new entry to The Galactic Blog, Galaxy hearts GCP.
- At least 11 Open positions in six countries on two continents.
- Updates to training materials.
- New releases.
- And some other news too.
If you have anything to add to next month’s newsletter, then please send it to outreach@galaxyproject.org.
GCC2019 Registration
Registration for the 2019 Galaxy Community Conference (GCC2019) is open! Discounted registration rates are available until 17MAY19.
GCC2019 will be held 1-6 July in Freiburg, Germany. The tenth GCC will have many familiar features from earlier years, including accepted and lightning talks, posters and demos, birds-of-a-feather gatherings (BoFs), training, and a CollaborationFest. GCC2019 brings the most significant conference program update in several years:
- some training will be integrated with the main conference,
- three days of conference instead of two,
- several parallel sessions,
- and a strong emphasis on organizing content by domain.
GCC2019, like every GCC before it, will be built around community. Training topics are nominated, selected, and presented by the community. Presentations will feature the full spectrum of Galaxy applications, enhancements and deployments from the community as well.
If you are working in data intensive life science research then there will not be a better place to share your work, learn from others, and find new collaborators.
Please find additional information on GCC2019 here.
Poster & Demo Abstract Submission is Still Open
The talk submission abstract deadline was last week, but poster and demo submissions remain open! Got something to show? Submit an abstract.
Fellowships: 2nd Round Due this Friday
Travel fellowships to attend GCC2019 are available. If you are interested, apply by this Friday, 10 May.
Sponsors
We are delighted to have GigaScience as a returning sponsor:
GigaScience
GigaScience is an online open access, open data, open peer-review journal published by Oxford University Press and BGI. The journal offers ‘big data’ research from the life and biomedical sciences, and on top of ‘Omics research includes the growing range of work that uses difficult-to-access large-scale data, such as imaging, neuroscience, ecology, systems biology, and other new types of shareable data. GigaScience is unique in the publishing industry as it publishes all research objects (data, software tools, source code, workflows, containers and other elements related to the work underpinning the findings in the article). Promoting Open Science, all published software needs to be under an OSI-license, all supporting data must be available and open, and all peer review is carried out transparently. Presenting workflows via our GigaGalaxy.net server, novel work presented at the meeting utilising Galaxy is eligible to a 15% APC if it is submitted to our Galaxy series.
GTN CoFest and community call
The Galaxy Training Network has developed an infrastructure to deliver interactive training based on Galaxy: one central place to aggregate training materials covering many current research topics. Each topic is supported by different tutorials developed and maintained by the community via a GitHub repository.

The Galaxy Training Network is organizing regular online CoFests (Collaboration/Contribution Fests) every 3 months on the 3rd Thursday for a day of the collaborative work on the training content.
The next one will be on the 16th of May. Anyone who would like to contribute on any other topics is very welcome to join.
We will coordinate via Hangout (drop-in channel we will keep open the whole day) and the GTN Matrix channel.
During the day, we plan to have GTN community calls to discuss the recent changes and future directions. To accommodate different time zones and schedules, we will have 2 calls (with similar discussions) at 8am (UTC + 2) and 4pm (UTC + 2) via Hangout.
If you want to join to help or to talk and learn about the project, let us know on the Matrix channel or by leaving a comment on the issue on GitHub
Upcoming events
These and other Galaxy related events are coming up:
Publications
265 new publications referencing, using, extending, and implementing Galaxy were added to the Galaxy Publication Library in the last month, pushing the total number of pubs past 7500.
Highlighted Publications
There were 15 Galactic and Stellar publications added. Here are the open access ones.
- Tackling “Big Data” with Biology Undergrads: A Simple RNA-seq Data Analysis Tutorial Using Galaxy, Matthew Anthony Escobar, William Morgan, Irina Makarevitch, Sabrina Robertson. CourseSource. https://doi.org/10.24918/cs.2019.13
- Containerization of Galaxy Workflows increases Reproducibility, Felix Bartusch, Maximilian Hanussek, Jens Krüger. Proceedings of the 4th bwHPC Symposium
- Desarrollo y despliegue de un workflow para el análisis genómico de Campylobacter Jejuni, José Antonio Barbero Aparicio, thesis, Universidad Autónoma de Madrid
- NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis, Guangyan Zhou, Othman Soufan, Jessica Ewald, Robert E W Hancock, Niladri Basu, Jianguo Xia. Nucleic Acids Research, gkz240, doi: 10.1093/nar/gkz240
- Comparative Dissection of Three Giant Genomes: Allium cepa, Allium sativum, and Allium ursinum, Vratislav Peška, Terezie Mandáková, Veronika Ihradská, Jiří Fajkus. Int. J. Mol. Sci. 2019, 20, 733.
- A rapid and simple method for assessing and representing genome sequence relatedness, bioRxiv, M Briand, M Bouzid, G Hunault, M Legeay, M Fischer-Le Saux, M Barret, doi: 10.1101/569640
- Identification of genetic relationships and subspecies signatures in Xylella fastidiosa, Nicolas Denancé, Martial Briand, Romain Gaborieau, Sylvain Gaillard, Marie-Agnès Jacques. BMC Genomics 201920:239, doi: 10.1186/s12864-019-5565-9
- Genotyping by sequencing can reveal the complex mosaic genomes in gene pools resulting from reticulate evolution: a case study in diploid and polyploid citrus, Dalel Ahmed, Aurore Comte, Franck Curk, Gilles Costantino, François Luro, Alexis Dereeper, Pierre Mournet, Yann Froelicher, Patrick Ollitrault. * Annals of Botany*, mcz029, doi: 10.1093/aob/mcz029. Published: 29 March 2019
- Genomic Variants Among Threatened Acropora Corals, Sheila A. Kitchen, Aakrosh Ratan, Oscar C. Bedoya-Reina, Richard Burhans, Nicole D. Fogarty, Webb Miller, Iliana B. Baums. G3: Genes, Genomes, Genetics, Early online March 26, 2019; doi: 10.1534/g3.119.400125
Publication Topics
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 162 | +Methods | 66 | +UsePublic | 36 | +Workbench | 26 | +RefPublic | |||
| 19 | +UseMain | 18 | +Reproducibility | 13 | +UseLocal | 9 | +IsGalaxy | |||
| 8 | +Cloud | 6 | +Shared | 5 | +HowTo | 5 | +Tools | |||
| 4 | +Other | 2 | +Education | 1 | +Unknown |
Galaxy Platforms News
The Galaxy Platform Directory lists resources for easily running your analysis on Galaxy, including publicly available servers, cloud services, and containers and VMs that run Galaxy. A lot was added in March:
Single Cell Omics Galaxy workbench
The Single Cell Omics workbench is a comprehensive set of analysis tools and consolidated workflows for single cell analysis. The current implementation comprises more than 20 bioinformatics tools dedicated to different research areas of single cell biology.
NetworkAnalyst
NetworkAnalyst is an online analysis environment for transcriptome profiling, network analysis, and meta analysis for gene expression data. NetworkAnalyst has developed and made available a public Galaxy server to support raw RNAseq processing for 17 species covering mammals, birds, bacteria, plants and parasites.
CellOrganizer
CellOrganizer creates generative models of the spatial organization of cells from microscope images and automatically provides geometries for spatial simulations of cell processes and behaviors. A Galaxy server provides access to CellOrganizer tools for users who do not have the resources to run the Matlab or Docker versions.
CIRM-CFBP
International Center for Microbial Resources - Plant Associated Bacteria (CIRM-CFBP) is providing web based access to tools (SkIf and KI-S) of IRHS-EmerSys lab on the CIRM-CFBP Galaxy instance.
Galaxy for Compound Activity Classification
Galaxy for Compound Activity Classification (GCAC)is a one stop solution for data generation, data preparation, feature selection, model building, prediction and retrieval of potential molecules. GCAC is available as a public Galaxy server, a VM, and an installable Galaxy tool suite from the toolshed.
MGS-FAST
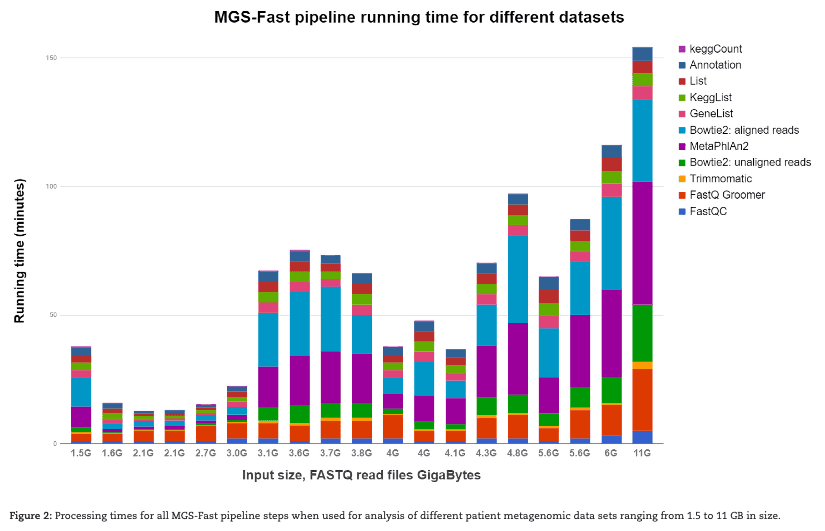
MGS-Fast is an analysis approach to rapidly map a metagenomics shotgun sequencing (MGS) data set and assign known functions to reads originating from microbial genes, producing counts of KEGG gene orthologs as output. This rapid annotation method is freely available as a Galaxy workflow within a Docker image.
Galaxy Platforms in Publications
We tag papers that use, mention, implement or extend public Galaxy platforms (servers, services, clouds, containers…). Here are the platforms that were referenced at least twice in the past month’s publications:
| # | Tag | # | Tag | # | Tag | # | Tag | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 19 | >Huttenhower | 11 | >ARGs-OAP | 9 | >RepeatExplorer | 7 | >Workflow4Metabolomics | |||
| 5 | >Galaxy-P | 4 | >Orione | 4 | >UseGalaxy.eu | 4 | >UseGalaxy.org.au | |||
| 3 | >epiGeEC | 2 | >APOSTL | 2 | >ARGalaxy | 2 | >CIRM-CFBP | |||
| 2 | >deepTools | 2 | >Pasteur | 2 | >SymD |
New Galactic Blog Post
This month we have a Galactic Blog post where Enis Afgan and Vahid Jalili describe recent efforts to Build up support for the Google Cloud Platform in Galaxy.
Who’s Hiring

The dark energy of irreproducible research is threatening the science universe! Please help the Galaxy push it back!
- Post-doc position in Computational Biology / Genomics / Transcriptomics / Bioinformatics, ETH Zurich, Switzerland
- Senior IT Systems Administrator, Rothamsted Research, Harpenden, United Kingdom
- Experienced Galaxy user / programmer / administrator for consulting work, Sarepta Therapeutics
- Genome Stability of Adult Stem Cells, Institut Curie, Dept. of Genetics and Developmental Biology, Paris, France
- Scientist - Molecular R&D, MilliporeSigma, Rockville, Maryland, United States
- Cloud Architect, Fred Hutchinson Cancer Research Center, Seattle, Washington, United States.
- ELIXIR Belgium has three Galaxy-related openings in Ghent:
- ELIXIR Open Science Community Manager, VIB-UGent Center for Plant Systems Biology
- ELIXIR Software developer data management tools, VIB-UGent Center for Plant Systems Biology
- ELIXIR Scientific Cloud Coordinator, VIB-UGent Center for Plant Systems Biology
- Software Developer, Harvard T.H. Chan School of Public Health, Boston, Massachusetts, United States. “Basic automated analysis workflows using Galaxy for 16S marker gene, metagenomic, and metatranscriptomic data leveraging existing software.”
- The The European Galaxy Team has open positions, Freiburg, Germany
- Software engineer, system analysts/administrators, data analyst, and a community and/or research manager
Have a Galaxy-related opening? Send it to outreach@galaxyproject.org and we’ll put it in the Galaxy News feed and include it in next month’s update.
Doc, Hub, and Training Updates
Updates from the Galaxy Training Materials in April:
Hub
The User Authentication and Authorization docs explain all of Galaxy’s features that (a) enable users to login to Galaxy using their social and institutional identities (e.g., using their Google account), and (b) allows them to securely authorize Galaxy to access their private cloud-based resources. Both (a) and (b) are described for admins (how to configure Galaxy for that feature) and users (how to use that feature) separately. Many thanks to Vahid Jalili, Kjell Petersen, and Enis Afgan for their contributions.
Also see Vahid Jalili’s slides on User Authentication and Cloud Authorization in the Galaxy project
GTN Training Materials
- New Mass spectrometry imaging: Finding differential analytes, by Melanie Foell
- New Creating a new tutorial - Writing content in Markdown tutorial by Bérénice Batut, Björn Grüning, Saskia Hiltemann, and Helena Rasche.
- New Name tags for following complex histories tutorial by Helena Rasche.
- New Bibliography feature for tutorials by Saskia Hiltemann and Helena Rasche.
- New RNA-Seq reads to counts tutorial by Maria Doyle.
- New RNA-seq counts to genes tutorial by Maria Doyle.
- New RNA-seq genes to pathwaystutorial by Maria Doyle.
- Update to Understanding UMIs tutorial by Mehmet Tekman.
ToolShed Contributions
 ](http://toolshed.g2.bx.psu.edu/)
](http://toolshed.g2.bx.psu.edu/)We didn’t get to this this month. Look forward to a double dose of tools next month.
Releases
New additions to the Galaxy Ecosystem.
Galaxy-parsec 1.12.1
Parsec is a set of automatically generated wrappers for BioBlend functions. I found myself writing a large number of small / one-off scripts that invoked simple bioblend functions. These scripts were impossible to compose and use in a linux-friendly manner. I copied and pasted code between all of these utility scripts.
Parsec is the answer to all of these problems. It extracts all of the individual functions I was writing as separate CLI commands that can be piped together, run in parallel, etc.
Pulsar 0.9.1
Pulsar is a Python server application that allows a Galaxy server to run jobs on remote systems (including Windows) without requiring a shared mounted file systems. Unlike traditional Galaxy job runners - input files, scripts, and config files may be transferred to the remote system, the job is executed, and the results are transferred back to the Galaxy server - eliminating the need for a shared file system.
galaxy-lib 19.5.2
galaxy-lib is a subset of the Galaxy core code base designed to be used as a library. This subset has minimal dependencies and should be Python 3 compatible. It’s available from GitHub and PyPi.
Other News
- The ELIXIR Galaxy Community held a training and tools workshop in Roscoff, France. Check out the meeting summary.
- SGCI Consultation Allows CloudLaunch to Benefit Researchers Across a Wide Variety of Fields, by Nayiri Mullinix
- Galaxy-ML: a GitHub repo from Qiang Gu of OHSU, providing a set of Galaxy tools wrapped with cutting-edge machine learning algorithms.